UCSF ChimeraX
UCSF ChimeraX (or simply ChimeraX)
is the next-generation molecular visualization program from the
Resource for Biocomputing,
Visualization, and Informatics (RBVI),
following UCSF Chimera.
ChimeraX can be downloaded free of charge
for academic, government, nonprofit, and personal use.
Commercial users, please see
ChimeraX commercial licensing.
ChimeraX is developed with support from National Institutes of Health R01-GM129325.
 ChimeraX on Bluesky:
@chimerax.ucsf.edu
ChimeraX on Bluesky:
@chimerax.ucsf.edu
Different representations of nucleotides can be shown with the
nucleotides
command or Toolbar
icons. Options include filled rings, slabs for bases
(box, muffler, or ellipsoid shape), bumps on slabs to show base orientation,
simple tubes instead of ribose atoms, and continuous or broken ladder rungs.
Nucleotide representations can be the same color as the ribbon
or a different color, and multiple nucleotide styles can be used
within a single structure.
See also: Presets menu
More features...
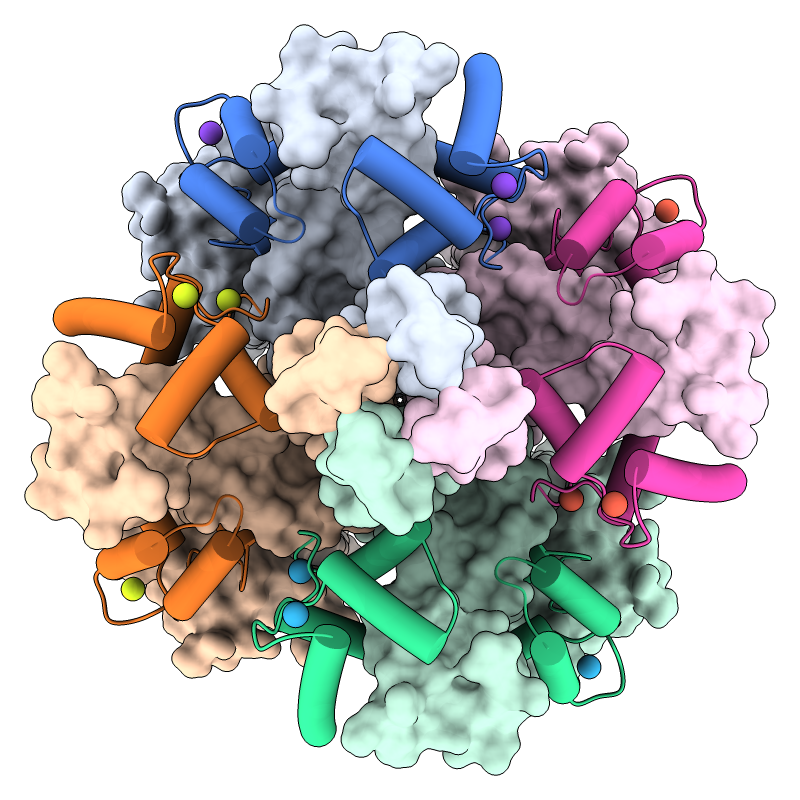
KCNQ1 is the pore-forming subunit of a cardiac potassium channel.
It binds to calmodulin, and mutations in either of these proteins
can cause congenital long QT syndrome, a dangerous
propensity for irregular heartbeats.
In the image, a structure of the KCNQ1/calmodulin complex
(PDB 5vms)
has been assembled into the native tetrameric form with the
sym command.
The view is from the cytoplasmic side, with
KCNQ1 shown as surfaces, calmodulin as cartoons, and calcium ions as balls.
A pastel palette
from ColorBrewer
has been used to color the surfaces, darkened with
color modify
for the cartoons, and “rotated” 45° in hue for the ions.
See the command file colormod.cxc.
More images...