Google™ Search
December 25, 2025
The RBVI wishes you a safe and happy holiday season!
See our
2025 card and the
gallery of previous cards back to 1985.
September 22, 2025
Mac users may wish to defer upgrading to MacOS Tahoe.
Currently on that OS the Chimera graphics window is shifted so that it covers
the command and status lines.
March 6, 2025
Chimera production release 1.19 is now available,
fixing the ability to fetch structures from the PDB
(1.19 release notes).
Previous news...
Please note that
UCSF Chimera is legacy software that is no longer being developed or supported.
Users are strongly encouraged to try
UCSF ChimeraX, which is under active development.

UCSF Chimera is a program for the interactive visualization
and analysis of molecular structures and related data,
including density maps, trajectories, and sequence alignments.
It is available free of charge for noncommercial use.
Commercial users, please see
Chimera commercial licensing.
We encourage Chimera users to try ChimeraX
for much better performance with large structures, as well as other major
advantages
and completely new features in addition to nearly all the capabilities
of Chimera (details...).
Chimera is no longer under active development.
Chimera development was supported by a grant from the
National Institutes of Health (P41-GM103311)
that ended in 2018.
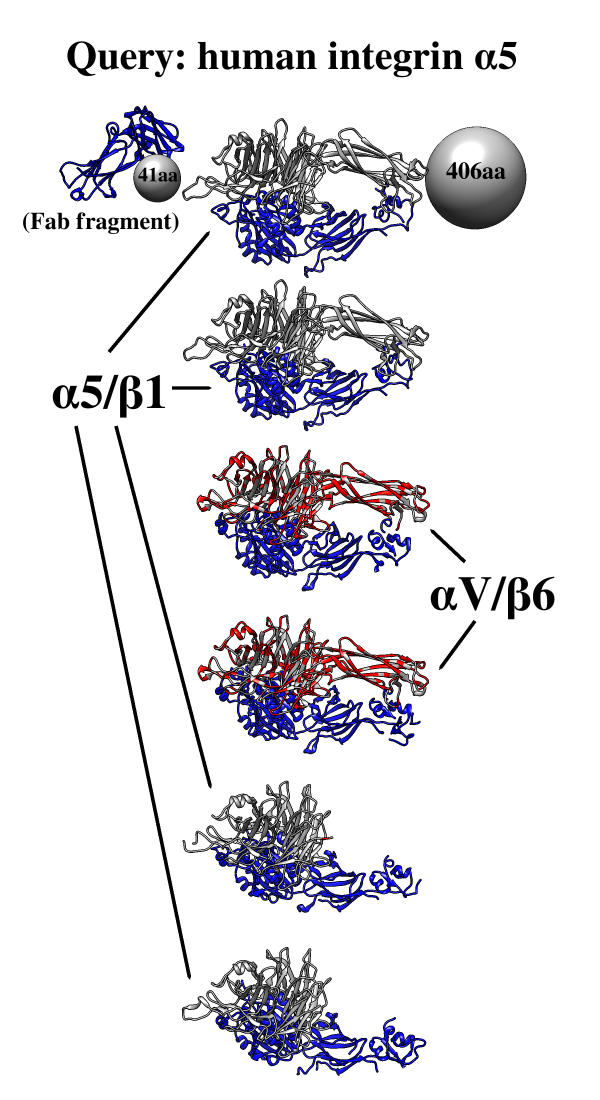
One use of
Multidomain
Assembler is to set up comparative modeling
and concatenation of existing structures to generate a full-length model
of a multidomain protein.
However, even without model-building, the byproduct is also useful:
a visual summary of the structures available for a query sequence,
optionally filtered by criteria such as BLAST score and % identity,
laid out horizontally in approximate N→C order relative to the query.
Overlapping hits are stacked vertically,
and segments without structural coverage are indicated with spheres.
By default, the multiple sequence alignment of the hits to the query
is also displayed.
The figure shows the results of command:
mda p08648 ~/Desktop/MDA limit 4 percent 50
with sequence mismatches in red and molecules other than the hit
chains in blue. Text and pointers have been added with
2D
Labels.
Multidomain Assembler is described in a
paper.
(More features...)
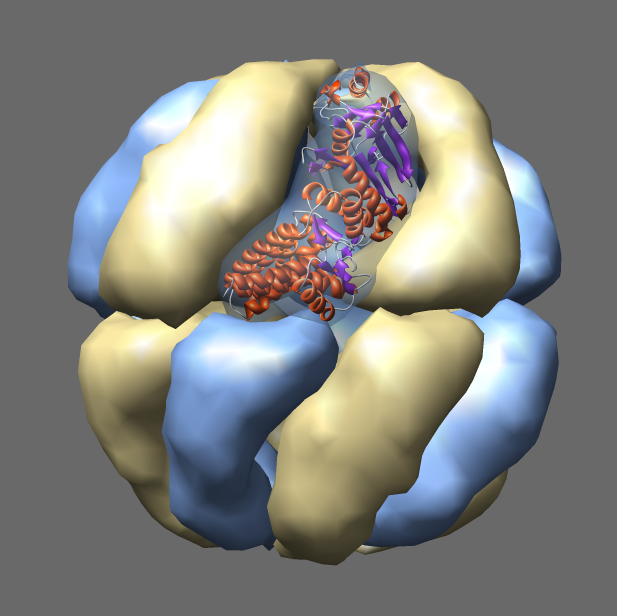
Thermosomes are hollow balls inside which proteins are folded.
They are found in the cytosol
of eukaryotes and in archaea. Eukaryotic thermosomes have 8 different
protein subunits, while archaeal ones are composed of one, two or three
different proteins. The one shown from Thermoplasma acidophilum has
two distinct proteins colored blue and yellow, each present in 8 copies.
The two proteins have 60% sequence identity and are very similar in
structure. One monomer is shown as a ribbon. Actin and tubulin are folded
by eukaryotic thermosomes.
Protein Data Bank model
1a6d.
(More samples...)
About RBVI
| Projects
| People
| Publications
| Resources
| Visit Us
Copyright 2018 Regents of the University of California.
All rights reserved.



