The sequence subcommands perform actions related to sequences in the Sequence Viewer or Profile Grid and associated structures:
The Tools menu can also be used to show the sequence of a structure chain in the Sequence Viewer. All of the other actions can be accessed from the Sequence Viewer context menu.
Usage: sequence chain chain-spec [ viewer true | false ]
The command sequence chain shows the sequence of the specified
biopolymer chain in the
Sequence Viewer,
although the graphical interface can be suppressed with viewer false
(for example, to run a script that uses the sequence data but not its display).
With viewer false, the alignment ID, sequence name,
and sequence are shown in the Log.
Only one structure chain should be specified per command, unless all
of the specified chains have exactly the same sequence.
See also:
Molecule Display icon
![]()
Other ways to start the Sequence Viewer: Independent of structure, sequence alignments and individual sequences can also be opened from files or fetched from UniProt. Other tools or commands may generate new sequence alignments (e.g., Blast Protein results, Matchmaker, sequence realignment).
Usage: sequence associate chain-spec [ alignment-ID:sequence-ID ]
Usage: sequence ( dissociate | disassociate ) chain-spec alignment-ID
Sequence-structure association (such as for synchronized selection) occurs automatically, but the commands sequence associate and sequence dissociate (same as sequence disassociate) allow more precise control, for example, of which structure chains are used for header calculations, or forcing or removing associations regardless of whether the number of mismatches would be tolerated by the automatic procedure.
The command sequence associate associates one or more structure chains with a sequence. The target sequence for association is specified by alignment ID, as reported in the title bar of the Sequence Viewer window, and the name or index number of the target sequence in the alignment, in the form: alignment-ID:sequence-ID (details...).
Alternatively, the sequence-ID can be omitted to associate each specified structure chain with the the best-matching sequence in the alignment. The alignment-ID can be omitted if only one alignment is open, or if the sequence-ID is also omitted; in the latter case, each specified structure chain will be associated with the best-matching sequence in each open alignment. If either or both are omitted, the colon (:) should also be omitted except in rare cases to disambiguate an alignment and sequence that have the same name.
For sequence dissociate, only the alignment needs to be specified, not an individual sequence, because a structure chain can only be associated with one sequence per alignment.
Usage: sequence match [ alignment-ID ] matchchain(s) to refchain [ cutoffDistance cutoff | none ] [ conservation percent ] [ columns list-of-positions ]
Superimpose the specified structures based on the alignment of their associated sequences. The alignment-ID is shown in the title bar of the Sequence Viewer window. (Conversely, to create a sequence alignment from a structural superposition, see msa3d.)
By default (cutoffDistance option not used), structures are fit iteratively with a cutoff of 2.0 Å for omitting farther-apart pairs from the fit. Specifying the cutoff as none turns iteration off. When iteration is performed, the sequence alignment is not changed, but residue pairs in the alignment can be pruned from the “match list” used to superimpose the structures. Fitting uses one atom per residue. In each cycle of fitting, pairs of atoms are removed from the match list and the remaining pairs are fitted until no matched pair is more than cutoff apart (default 2.0 Å). The atom pairs removed are either the 10% farthest apart of all pairs or the 50% farthest apart of all pairs exceeding the cutoff, whichever is the lesser number of pairs. The result of iteration is that the best-matching “core” regions are maximally superimposed, and conformationally dissimilar regions such as flexible loops are not included in the final fit, even though they may be aligned in the sequence alignment.
The conservation option specifies omitting columns with less than percent identity from the initial fit.
The columns option specifies omitting columns other than those listed from the initial fit. The list must consist of positive integers separated by commas only; hyphenated ranges cannot be used.
The number of atom pairs fitted and the resulting RMSD are reported in the Log and the status line at the bottom of the ChimeraX window. See also: matchmaker, align
To create a sequence alignment from a structural superposition, see msa3d.
Usage: sequence expandsel [ alignment-ID ]
Expand the current selection to all residues associated with the same column(s) of the specified sequence alignment, or if none is specified, all sequence alignments. The alignment-ID is shown in the title bar of the Sequence Viewer window.
Usage: sequence rename [ alignment-ID:sequence-ID ] new-name
Rename the specified sequence. The sequence is specified by its alignment ID, shown in the title bar of the Sequence Viewer window, plus the sequence's name or index number in the alignment, in the form: alignment-ID:sequence-ID (details...). The new-name cannot include a colon (:) and should be enclosed in quotation marks if it includes any spaces.
Usage: sequence header [ alignment-ID ] header-name ( show | hide | save filename )
The command sequence header shows, hides, or saves a sequence header to a file. (It can also be used to change the sequence Headers preferences, but command details are omitted here because normally the Settings dialog will be used instead.)
The header-name can be consensus, conservation, or rmsd, although there will only be an effect when that header is available (for example, an RMSD header is only available for alignments associated with at least two structures). Headers are saved to a simple text format that lists the alignment positions and values. The filename can be given as a pathname or the word browse to bring up a file browser window for choosing the name and location interactively. If multiple alignments are open but an alignment-ID is not specified, showing/hiding affects all applicable alignments. However, saving only works for a single header at a time, so an alignment-ID must be given when more than one alignment is open.
Usage: sequence refseq [ alignment-ID:sequence-ID ]
The overall numbering of an alignment is shown across the top at the first position and every tenth position thereafter. By default, there is no reference sequence, and the alignment is simply numbered 1, 11, 21, etc., regardless of whether the columns contain gaps. However, with sequence refseq, any of the individual sequences can be specified as the reference for numbering instead. The sequence to use is specified by its alignment ID, shown in the title bar of the Sequence Viewer window, plus the sequence's name or index number in the alignment, in the form: alignment-ID:sequence-ID (details...).
When the reference sequence has a gap character at a position where a number is to be shown (first column, eleventh column, etc.), how that position falls relative to the reference is shown in parentheses. For example, “(<1)” would be shown for alignment positions before the reference's first residue, and “(43/44)” for positions that fall between residues 43 and 44 of the reference.
Omitting the sequence identifier returns to the default numbering:
Usage: sequence refseq [ alignment-ID ]
If more than one Sequence Viewer window is present, the alignment-ID is required. The alignment-ID is shown in the title bar of the Sequence Viewer window.
Usage: sequence identity alignment-ID [ denominator shorter | longer | nongap ]
Usage: sequence identity alignment-ID alignment-ID:sequence-ID [ denominator shorter | longer | nongap ]
Usage: sequence identity alignment-ID:sequence-ID alignment-ID:sequence-ID [ denominator shorter | longer | nongap ]
The sequence identity command calculates the pairwise percent identity between sequences of the same length (including gaps, as shown in the Sequence Viewer window). The calculation is always pairwise, but can be performed for all-by-all pairs within a single alignment, or all-by-one, or between two specific sequences. An entire alignment is specified by its ID, shown in the title bar of the Sequence Viewer window, and an individual sequence by the alignment ID plus the sequence's name or index number in the alignment, in the form: alignment-ID:sequence-ID (details...).
Results are listed in the Log. For each pair, the number of columns with identical residues is given as a percentage of the specified denominator:
Usage: sequence refreshAttrs alignment-ID
For structure chains associated with more than one sequence alignment, make sure that the current attribute values are consistent with a specific alignment. The alignment-ID is shown in the title bar of the Sequence Viewer window.
Usage: sequence search chain-spec [ database pdb | afdb ] [ showTable true | false ] [ evalueCutoff max-evalue ] [ identityCutoff min-pid ] [ maxHits N ] [ trim true | false | chains,sequence,ligands ] [ alignmentCutoffDistance dist ] [ saveDirectory results-folder ]
The sequence search command performs a fast MMseqs2 search of the PDB or AlphaFold Database with a protein structure chain as the query, using the RCSB web service. The Foldseek (Similar Structures) tool provides a graphical interface to running this command. See also: similarstructures, foldseek, blastprotein
MMseqs2 (Many-against-Many sequence searching) is described in:MMseqs2 enables sensitive protein sequence searching for the analysis of massive data sets. Steinegger M, Söding J. Nat Biotechnol. 2017 Nov;35(11):1026-1028.
MMseqs2 desktop and local web server app for fast, interactive sequence searches. Mirdita M, Steinegger M, Söding J. Bioinformatics. 2019 Aug 15;35(16):2856-2858.
The showTable option (default true) indicates whether to show the results in the Similar Structures tool, which facilitates exploring large sets of protein structures by efficiently showing them in 3D as backbone traces and in 2D as sequence alignment schematics or scatter plots based on conformation. These analyses can also be performed with the similarstructures command by referring to a set of results by name. The name assigned to the results (mm1 or mm2) is reported in the Log when the search is run.
Choices of database to search:
The hits can be limited with the following options:
The trim and alignmentCutoffDistance options do not act immediately. Instead, they specify how to process the hit structures if later opened from the Similar Structures tool or with the command similarstructures open. The structures are fetched from the respective database (Protein Data Bank or AlphaFold Database) and processed as follows:
Search results are saved in a similar structures file (suffix .sms, a JSON file format specific to ChimeraX) with filename based on the query name and the database searched. The file will be listed in the File History for easy access, and simply opening it loads the set of results into the Similar Structures interface. The saveDirectory option allows specifying the save location, either directly or as the word browse to specify it interactively in a file browser window (default location ~/Downloads/ChimeraX/MMseqs2/).
Usage: sequence update chain-spec [ alignment alignment-ID ]
Standard PDB files have SEQRES records containing the sequence of each biopolymer chain in the structure, and similarly, standard mmCIF files include this information in specific tables. In many structures, some of the residues in the sequence are missing from the atomic coordinates because their positions could not be resolved from the experimental density (due to disorder, etc.). However, atomic coordinate files output from various modeling programs often lack sequence information, so that the sequence is only inferred from the coordinates. The sequence update command is useful for adding full sequence information to such structures. For example, the full sequence could be opened from a FASTA file or by fetching it from UniProt, associated with the chain, and then used to update the chain's sequence information. This command only applies to the sequences associated with the specified chains, limited to a specific alignment as needed. The alignment-ID is shown in the title bar of the Sequence Viewer window. The sequence must cover all of the residues in the structure chain.
Usage: sequence align alignment-ID [ replace true | false ] [ program clustalOmega | muscle ]
Usage: sequence align chain-spec [ program clustalOmega | muscle ]
Usage: sequence align sequence1,sequence2[,sequence3...,sequenceN] [ program clustalOmega | muscle ]
The sequence align command calculates a new alignment of the specified protein sequences using a web service hosted by the UCSF RBVI. (To create a sequence alignment from a structural superposition instead, see msa3d.) The result is opened in a new Sequence Viewer window, except that replace true (default false) can be used to specify overwriting an existing alignment when all of its sequences are being realigned.
The sequences to align can be specified collectively by:
sequence align ldlr_rat,ldlr_mouse,ldlr_human
open myfile.msf
sequence align 1,2,3,-1
– OR (if multiple sequence windows are open) –
sequence align myfile.msf:1,myfile.msf:2,myfile.msf:3,myfile.msf:-1
The program can be either of two choices:
See the README file at the Clustal Omega website for details. Users should cite:
Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Sievers F, Wilm A, Dineen D, Gibson TJ, Karplus K, Li W, Lopez R, McWilliam H, Remmert M, Söding J, Thompson JD, Higgins DG. Mol Syst Biol. 2011 Oct 11;7:539.
Users should cite:
Muscle5: High-accuracy alignment ensembles enable unbiased assessments of sequence homology and phylogeny. Edgar RC. Nat Commun. 2022 Nov 15;13(1):6968.
Usage: sequence grid alignment-ID label residue-spec [ palette palette ] [ range low,high | full ] [ bgColor color-spec ] [ height h ] [ offset x,y,z ] [ onTop true | false ]
|
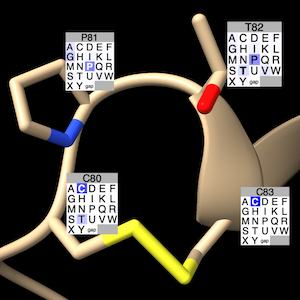
Label residues of structure chains associated with an alignment in Profile Grid with color-coded grids of residue-type prevalences. The alignment-ID can be specified if more than one Profile Grid is present. The label grids show the prevalences of the 20 standard amino acids and gap at the corresponding column in the alignment, by default coloring from white to blue with increasing prevalence:
palette 0,white:1,blue range 0.0,1.0
The structure residue's type is shown in the label title (along with its number) and with bold font in the grid. The bgColor is used behind the label text and for filling in empty cells in the last row of the label (default #B4B4B4
The label height h defaults to 1.5 Å. The offset x,y,z is the position of the lower left corner of the grid relative to the residue's α-carbon along X, Y, and Z in screen coordinates (default 0,0,3 Å). The onTop setting is whether to draw the labels on top of other objects regardless of their relative positions in 3D (default false). The labels can be removed with label delete.
See also the Profile Grid context menu.