This is a side example from the Images for Publication tutorial describing the creation of static, shadowed renderings with neon (not available on Windows).
| [images shown at 25%; click for full size] | |

|

|
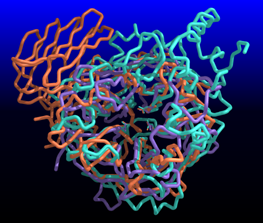
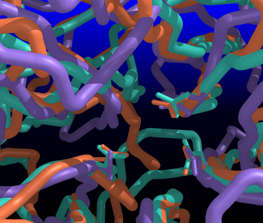
| Comparison of glycoside hydrolases from different families. Invertase (family 32) is coral, arabinanase A (family 43) is purple, and levansucrase (family 68) is turquoise. Despite differences in sequence, substrate, and catalytic mechanism, the enzymes have similar overall folds (left) and a set of highly conserved acidic residues in the active site (right). | |
The structures and positions are the same as in the main Images for Publication tutorial. Starting from the same Chimera session (images.py), the following commands were used:
Command: ~ribbonThe last command improves the interactive appearance but does not affect images saved with neon.
Command: sel @/display
Command: chain @ca
Command: ~disp ions
Command: disp sel
Command: ~sel
Command: repr stick
The neon command calls the external programs neon and conic. Parameters can be set within two separate input files, one for each program. Both of these input files were present in the working directory:
Command: reset overall(Depending on the system, this may take several minutes to complete.) The command-line flags are explained in the conic manual page; -c indicates the file with conic parameters, -x and -y indicate the pixel dimensions of the output image, -o indicates the output file name, and -s indicates that the image should be shown on the screen when the computation is complete. The -s flag works only on Linux, Mac, and SGI systems, and not all image formats can be displayed on all systems. On SGI systems, -f tiff would also be needed to specify TIFF output.
Command: neon -c conic.dat -x 1050 -y 890 -o neon1.tif -s
After the saved position closeup was restored and clipping planes were again adjusted to abut the structures, the same command (except for the output file name) was used to generate neon2.tif.