This multi-disciplinary research program, begun in 2000, focuses on the pharmacogenetics of membrane transport proteins that play a role in drug response pathways. The project seeks to understand the genetic basis for variation in drug response for drugs which interact with membrane transport proteins. This class of proteins is of great pharmacological importance, as it provides the target for about 30% of the most commonly used prescription drugs and is a major determinant of the absorption, distribution and elimination of many others. The project seeks to test the hypothesis that variations in the membrane transporter gene sequences underlie inter-individual differences in response to such drugs. Most human sequence variation is in the form of single-nucleotide polymorphisms (SNPs). The project evaluates the prevalence and distribution of both common and rare (<1% of the population) alleles by genotyping an ethnically diverse reference sample. Our program is structured around six interacting cores: genomics, cellular phenotyping and functional genomics, clinical phenotyping, biostatistics, shared resources/collaborations, and bioinformatics. The project also facilitates studies by contributing data to the publicly available knowledge base, PharmGKB (http://www.pharmgkb.org/). This project is funded by the National Institute of General Medical Sciences.
The specific aims of the PMT project are to:
- Identify sequence variants in coding and noncoding regions of genes encoding selected membrane transport proteins;
- Through integrated computational and experimental methods in cells, model organisms and in humans, functionally characterize sequence variants in membrane transporters that play a role in drug disposition and response;
- Determine the biological relevance of gentic variants in membrane transporters to clinical drug disposition and response in ethnically diverse human populations;
- Deposit the data in the Pharmacogenetics Network Database, PharmGKB.
The Resource for Biocomputing, Visualization, and Informatics (RBVI) plays a major role in the overall PMT project, acting as the bioinformatics core. The RBVI has developed methods for large-scale data collection, storage, analysis, presentation of experimental results (Figure 1), and uploading of these results to the PharmGKB database. Data processing begins when the Genomics Core uploads resequencing data via a web form on the secure PMT intranet site. The data undergo validation during upload and again during processing. Per-sample data is collated into observed snps, then computationally analyzed and characterized. Results are presented on the PMT project's web site, which the RBVI has created and maintains. Included in the data analysis is prediction and display of secondary structure, and indication of which amino acids and their location within the predicted secondary structure are affected by the experimentally-determined SNPs. Thus, this work deals with all aspects of the sequence-structure-function triad and meshes well with the goals of the RBVI Resource.
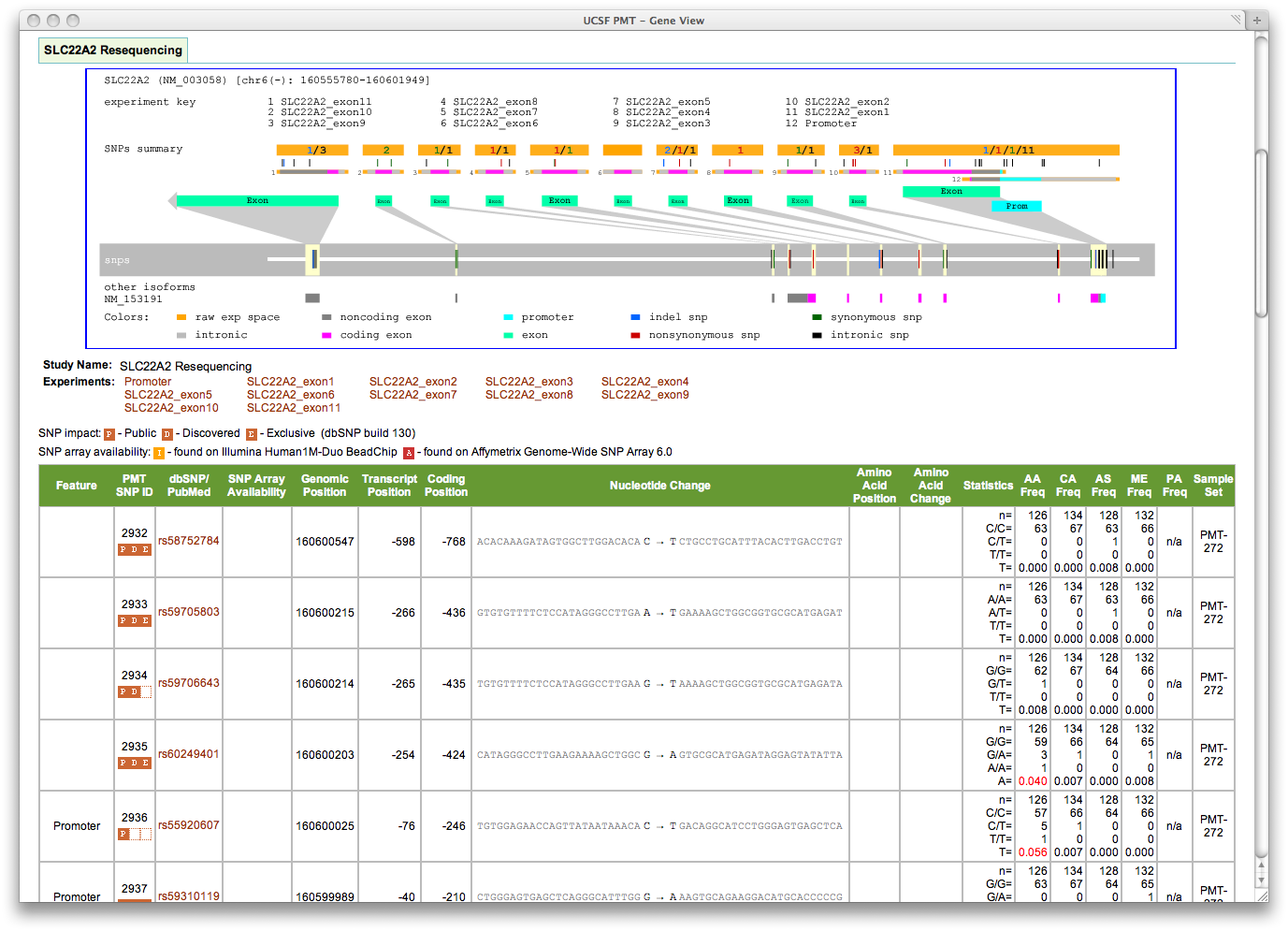
Figure 1: Screen shot from PMT web site showing results of data analysis for SLC22A2 transporter. Results are presented in both a graphical and tabular format. The graphic includes assay and snp information and provides genomic context. Assay regions are characterized as to gene features, such as intron or coding and noncoding exon. Snps are characterized by genomic position and affect on amino acid. Tabular data includes dbSNP identifier, genomic position, transcript and coding offsets, genomic context, genotype counts, and frequency by ethnic group. Links allow researchers to explore experimental details.
References:
- SNP Analysis and Presentation in the Pharmacogenetics of Membrane Transporters Project. D. Stryke, C.C. Huang, M. Kawamoto, S.J. Johns, E.J. Carlson, J.A. DeYoung, M.K. Leabman, I. Herkowitz, K.M. Giacomini, and T.E. Ferrin, Pac. Symp. Biocomput. 2003, pp. 535-547. Edited by R.B Altman, A.K Dunker, L. Hunter, T.A. Jung, and T.E Klein. World Scientific Publishing, 2003.
- Natural Variation in Membrane Transporter Genes Reveals Evolutionary and Functional Constraints. Maya K. Leabman, Conrad C. Huang, Joseph DeYoung, Elaine J. Carlson, Travis Taylor, Melanie de la Cruz, Susan J. Johns, Doug Stryke, Michiko Kawamoto, Deanna L. Kroetz, Thomas E. Ferrin, Andrew G. Clark, Neil Risch, Ira Herskowitz, and Kathleen M. Giacomini. Proc. Natl. Acad. Sci. USA, 100(10):5896-5901, 2003.
- Enhancing Data Sharing in Collaborative Research Projects with DASH. T.E. Ferrin, C.C. Huang, D.M. Greenblatt, D. Stryke, K.M. Giacomini, and J.H. Morris. Pac. Symp. Biocomput., pp. 260-271. Edited by R.B Altman et al., World Scientific Publishing, 2005.
