MatchMaker Results
The test cases involve structures with low pairwise sequence identity:
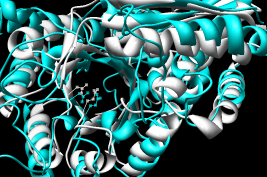
- Pairwise - 2mnr (mandelate racemase) 4enl (enolase)
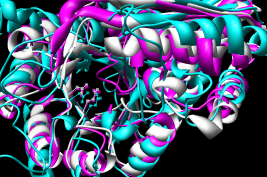
- Multiple - 2mnr 4enl 1nu5 (MLE II)
The
MatchMaker tool
in Chimera
constructs a sequence alignment and superimposes the structures accordingly.
As of version 1.2129 (June 2005), secondary structure can be used
to help construct the alignment, which allows pairs with lower
percent identity to be superimposed correctly. However, MatchMaker
does not handle/detect circular permutations. Results
from using MatchMaker with the default settings in version 1.2129:
Results are about as good as with the
superposition servers.
The multiple superposition here is not truly multiple, but
constructed from pairwise matches of 4enl and 1nu5 to the
reference structure, 2mnr.
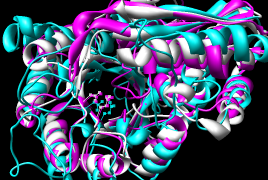
December 2005: Results are the same with Chimera version 1.2184 and the
newer MatchMaker defaults (30% weighting of secondary structure score):
Elaine Meng / meng@cgl.ucsf.edu /
home page