

Tom Goddard
September 2001
The helixhunter program can find alpha helices in a density map. Sequence analysis programs can predict the positions of helices in the amino acid sequence of the protein. To make a correspondence between the helices found in the density map and the sequence it is helpful to trace the protein backbone. A simple approach is to interactively place markers on the density map to draw chains connecting the helices in 3 dimensional space. This can be done using Chimera as shown in the figures.
The simulated density map for Bluetongue virus capsid protein was provided by Matthew Baker and Wen Jiang of the National Center for Macromolecular Imaging.
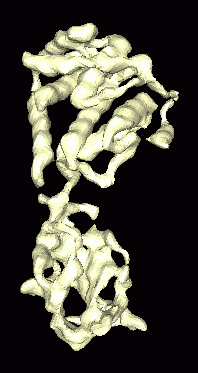
Figure 1: Isosurface of density map for Bluetongue virus capsid protein. Density map was simulated at 8A resolution from 1BVP crystal structure. |
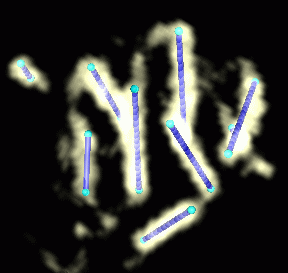
Figure 2: Helices are the most noticable features of the density map. These helices were created with the mouse by placing markers at the end points of prominent density sausages. Using helices predicted by helixhunter would eliminate this subjective choice.
|
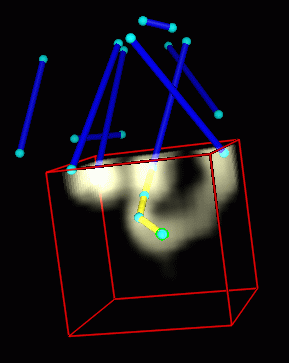
Figure 3: The turns are traced by pressing a mouse button to drop markers that define a chain from one helix to the next. Displaying only the region of the density map within 10 angstroms of the current marker simplifies the tracing. Fewer overlaps in the small region permit use of lower density thresholds. |
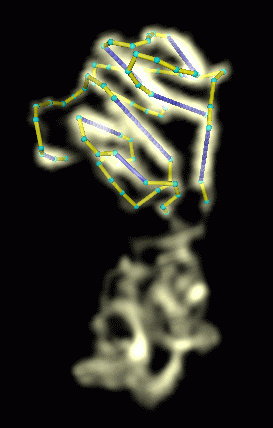
Figure 4: The alpha helical domain can be traced in about a half hour. Comparison with the crystal structure shows that the helices are joined in the correct order. |
Laboratory Overview | Research | Outreach & Training | Available Resources | Visitors Center | Search