The Research Collaboratory for Structural Bioinformatics-Protein Data Bank (RCSB-PDB) (Berman, 2000), in collaboration with world-wide PDB partners (www.wwpdb.org), currently houses more than 34,000 entries describing structures of biological macromolecules derived from X-ray crystal structures, NMR and cryo electron microscopy experiments. The mission of the RCSB-PDB is to provide the most accurate, well-annotated data about macromolecular structure in the most timely and efficient way possible to facilitate new discoveries and advances in science. In the past decade, the PDB has continually improved the data content and quality of deposited macromolecular structures. To handle the large volume of incoming deposits, the PDB has implemented a data pipeline (see diagram below) to efficiently accept, validate, archive and distribute user data. Validation directly affects the quality of the data content of the PDB. One of the main components of validation is visual examination of submitted data, and the PDB uses UCSF Chimera extensively for its molecular visualization needs. These needs have become particularly critical over the past several years with the steady increase in complex entries with high symmetry or large numbers of chains. Over the past few years we have benefited from a close collaboration with the Resource for Biocomputing, Visualization, and Informatics (RBVI), and we anticipate this collaboration will continue to evolve.
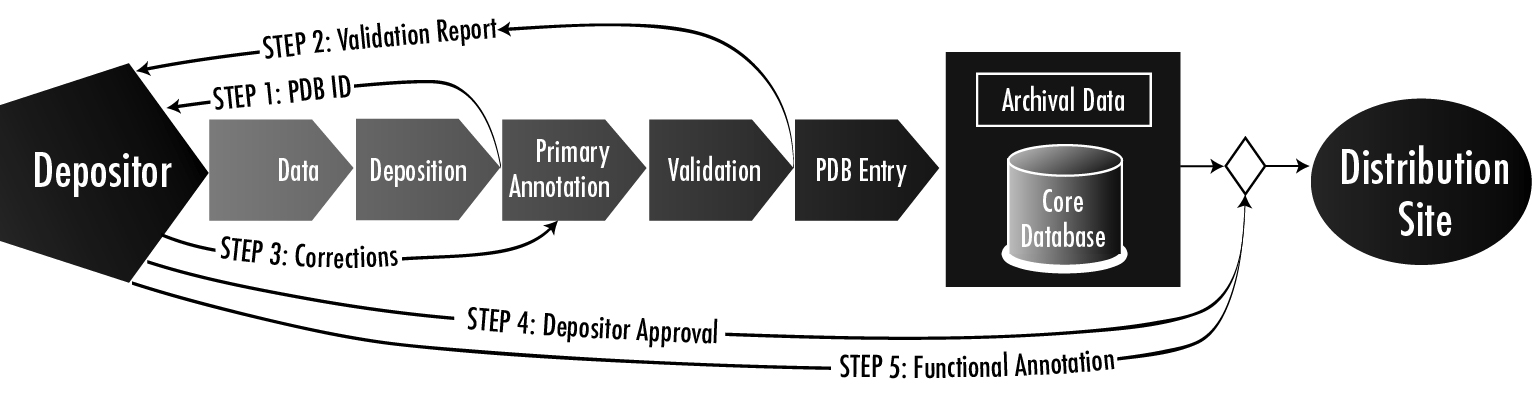
Figure caption: Details of the PDB data processing step of the overal data pipeline. An important part of the validation process is visual inspection of the data using Chimera.
The PDB and the services built on it are utilized by thousands of scientists worldwide including biologists, chemists and computer scientists who are seeking to interpret biological processes in molecular terms. Virtually every pharmaceutical company uses the contents of the PDB for drug discovery. The PDB also serves an educational resource for teachers and students from K-12 grades up to university levels.
Below we describe two areas in which our collaborations with the RBVI have benefited our research projects together with some of our future plans involving Chimera.
Promoting mmCIF as the Preferred Structure File Format The original PDB format was designed around an 80-column data format dating back to the days of batch processing "Hollerith" punch card technology. The fixed-width nature of the data fields in the original format specification has imposed a number of significant limitations for representing data; perhaps most notable is the limit of 99,999 atoms per structure, which is less than the number of atoms in some structures now available. In 1997, a replacement format, the macromolecular crystallographic information file (mmCIF), was proposed (Bourne, 1997). mmCIF has a number of advantages over the original PDB format, including a better machine-readable format, ability to handle much larger structures, and extensibility to accommodate new data types, just to name a few. A drawback to mmCIF is the complexity of its specification, which has discouraged some software developers from migrating to the new format despite the increasingly severe limitations imposed by the original PDB format. In particular, the number of molecular visualization applications that accept mmCIF files is very small, with Rasmol perhaps being the best known example.
The recent addition to Chimera of the capability of reading mmCIF formatted structure files, and the planned addition of also being able to write such files, will do much to promulgate the mmCIF standard and also provide a means to take advantage of many of the new features that mmCIF offers. For example, Chimera can utilize the biological unit descriptions in mmCIF files to generate and display biologically relevant multimeric molecules. (In the original PDB format, these descriptions are embedded in "REMARK" records that contain free-form text, and even though some depositors used a common data format convention, not all entries conform to this informal convention thereby making unit cell data unreliable for the end user.) The extensibility of Chimera also allows a convenient way to add new user interfaces to the system that allow for browsing through mmCIF data. Because mmCIF explicitly defines many more data fields, an mmCIF "data browser extension" to Chimera could take advantage of the semantics of these standard fields to present a more intelligent interface to the data for the scientist-user. Finally, adding the ability to validate and save mmCIF files will make Chimera a much more useful tool for data depositors, who can preprocess data using their favorite tools of choice and then use Chimera to prepare this data for submission to the PDB.
Validation of Virus Biological and Crystal Unit Cell Matrices
The majority of PDB entries describe crystals structures of
macromolecules in terms of their asymmetric units. An asymmetric unit
is the smallest portion of the crystal structure that can be used to
generate the unit cell through symmetry operators. Symmetry may also
be used to describe the biological complex that has been shown to be,
or is believed to be, functional. (See the PDB
biological unit tutotial
for a detailed description.)
The PDB currently houses more than two hundred virus structures. For these entries, the correctness of the symmetry description is particularly difficult to verify because few programs can display the large number of asymmetric units needed to generate the complete complex. We have adopted the MultiScale extension of Chimera as our primary means to identify errors in virus symmetry descriptions, because computationally it is much more efficient to apply symmetry operations to volumes rather than coordinates. Invalid symmetry descriptions are typically very easy to identify visually because the computed virus exhibits obvious gaps and clashes, while valid entries have a regular, non-overlapping appearance (see images).
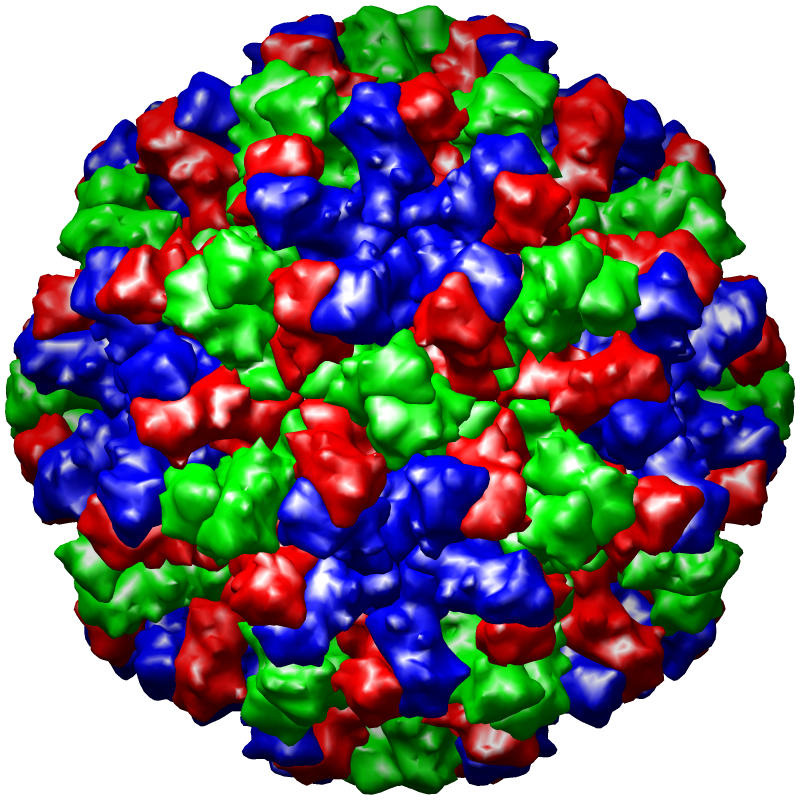
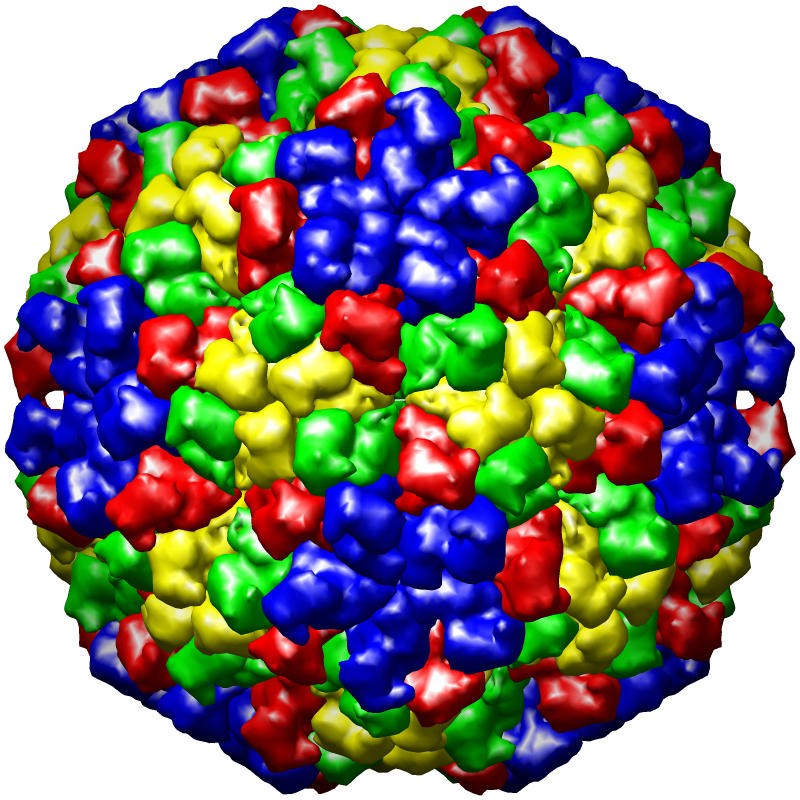
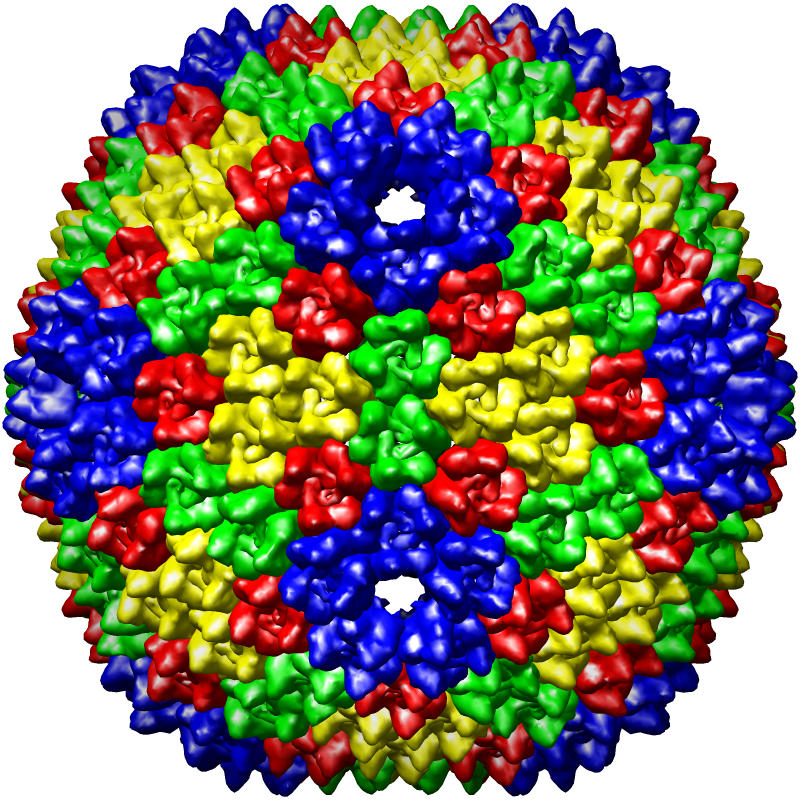



Figure caption: PDB virus structure entries rendered using the UCSF Chimera MultiScale extension tool. From left to right, the structures are of Dengue virus (1k4r), Nudaurelia Capensis ω Virus (1nwv), and Bacteriophage PRD1 (1gw7).
Chimera scripts have been developed by the RBVI as part of our collaboration for automating some procedures, such as creating images of batches of PDB entries for later perusal. A future Chimera extension for automatically identifying clashing regions has also been discussed. The development of these tools will help facilitate validation of new virus structure submissions.
References:
- Berman HM, Battistuz T, Bhat TN, Bluhm WF, Bourne PE, Burkhardt K, Feng Z, Gilliland GL, Iype L, Jain S, Fagan P, Marvin J, Padilla D, Ravichandran V, Schneider B, Thanki N, Weissig H, Westbrook JD, Zardecki C. The Protein Data Bank. Acta Crystallogr D Biol Crystallogr. 58(Pt 6 No 1):899-907, 2000.
- Tagari M, Newman R, Chagoyen M, Carazo JM, Henrick K. New electron microscopy database and deposition system. Trends Biochem Sci. 27(11):589, 2002. (PMID: 12417136)
- Bourne PE, Berman HM, McMahon B, Watenpaugh KD, Westbrook J, Fitzgerald PMD. The Macromolecular Crystallographic Information File (mmCIF). Methods in Enzymology. 277:571-590, 1997. (http://mmcif.pdb.org/pubs/methenz.html)
- Yan X., Chipman P.R., Battisti A.J., Bergoin M., Rossmann M.G. and Baker T.S. (2005). The Structure of the T=147 Iridovirus, CIV, at 13A Resolution. Proceedings of Microscopy and Microanalysis, 11(S02): 134-135.
