

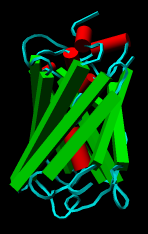
PipesAndPlanks creates a VRML representation of protein secondary structure in which helices are shown as "pipes" (cylinders) and strands are shown as "planks." Protein helix and strand assignments are taken from the input structure file or generated with ksdssp. Segments other than helices or strands are treated as turns (even though they may not be defined as such in the input file). PipesAndPlanks does not work for nonprotein molecules such as nucleic acids.
There are several ways to start PipesAndPlanks, a tool in the Depiction category. Starting PipesAndPlanks opens a dialog box. The model of interest should be chosen from the pulldown list of open molecule models (indicated by the black inverted triangle near the top of the dialog).
Helix color, Strand color, and Turn color assignments can be changed by clicking the respective color well. Helix radius, Strand width, Strand thickness, Turn width, and Turn thickness parameters are in angstroms. If Fixed [helix radius, strand width, or strand thickness] is false, the specified value is ignored and the parameter instead depends on the distances from the constituent alpha-carbons to the axis of the helix or strand. If the ratio of maximum to minimum distance from alpha-carbons to the axis exceeds the [Helix or Strand] split threshold (and the corresponding Split curved... option is true), the pipe or plank will be split into two parts, each with its own axis.
Clicking OK creates the display and dismisses the dialog; clicking Apply creates the display without dismissing the dialog. The pipes-and-planks VRML representation is placed in the same model number as the corresponding molecule. It can be closed or hidden independently of the corresponding molecule model with the Model Panel, or undisplayed/displayed (hidden/shown) with the command objdisplay. Close dismisses the dialog, and Help opens this manual page in a browser window.