February 27, 2007
Adds command transition for varying volume contour level, volume and molecule and molecular surface transparency, making multiscale surfaces fly away radially, and smoothly changing radius in icosahedral surface tool. These are for making animations.
Change contour level of model #0 from 3.0 to 1.5 for 100 frames.
transition contour_level #0 3.0 1.5 100
Fade out volume data contour surface over 100 frames.
transition transparency #0 0 1 100
Fade out chain A of molecule #0 over 100 frames.
transition transparency #0:.A 0 1 100
Fade MSMS molecular surface to 50% transparency over 100 frames.
transition msms_transparency #0 0 .5 100
Move protein surfaces from multiscale model with id #1 radially by a factor of 2 over 10 successive frames.
transition radial_motion #1 2 10
Reverse the radial preceding motion:
~transition radial_motion #1 2 10
Move only chains A, B, and C radially leaving the others in place:
transition radial_motion #1:.A,.B,.C 2 10
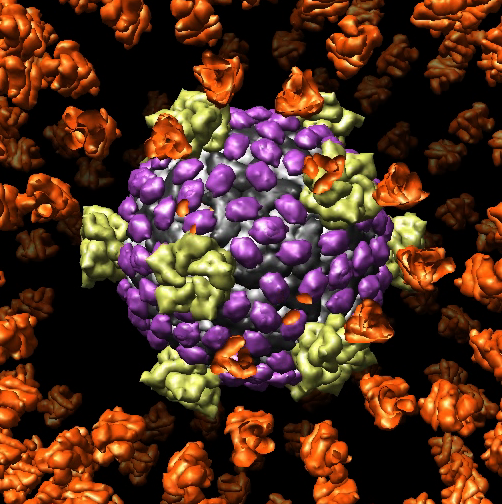
Here's a Chimera command script flyaway.cmd for successively removing layers of orthoreovirus virus (PDB 2cse) as shown in movie reovirus.mov.
roll y .25 10000 wait 200 transition radial_motion #1:.S,.D,.E,.F,.M,.N,.O,.G,.H,.I 4 400 wait 200 transition radial_motion #1:.A,.B,.C,.P,.Q,.R,.J,.K,.L,.T 4 400 wait 200 transition radial_motion #1:.X,.Y,.Z 4 400 wait 200 transition radial_motion #1:.U 4 400 wait 200 transition radial_motion #1:.V,.W 4 400 wait 200 transition radial_motion #1:.1 4 400 freeze
It can be run using the Chimera command open flyaway.cmd. The wait command delays a specified number of graphics frames before running the next command. The roll command spins about an axis and freeze stops the roll.
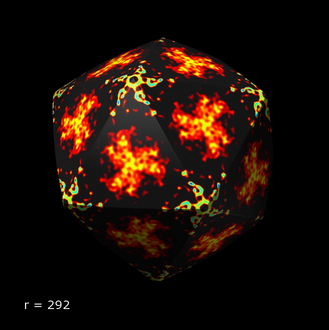

Increase icosahedron radius from 30 to 80 in 1 angstrom steps.
transition icosahedron_radius 30 80 1
The radius is shown in the lower left corner of the graphics window. Here is a movie showing the icosahedron colored according to values of a ricedwarf EM density map.
Use the Model Panel (menu entry Favorites / Model Panel) to find out the model numbers.
Chimera does not correctly render transparency when 2 or more models are transparent and overlapped. The transparent model with the higher model id number will appear in front of other transparent models regardless of its actual position in space.
Transparency values vary from 0 (totally opaque) to 1 (fully transparent). A fully transparent model is invisible. Technically the model is still displayed, so for instance it will be outlined in green if it is selected. In some applications it may be desirable to undisplay the model after fading it out completely.
Opening a volume data set causes it to be displayed and the Volume Viewer dialog is shown. The contour level reported in the volume dialog is not updated when the transition command is used.
The Multiscale Models tools (menu entry Tools / Higher-Order Structure / Multiscale Models) creates multiscale surfaces. The multiscale tool does not know that the proteins have been moved. Changing the multiscale surface resolution will restore the proteins to their original locations.
The Icosahedron Surface dialog is obtained from menu entry Tools / Higher-Order Structure / Icosahedron Surface.
transition contour_level <volume-surface-id> <start-level> <end-level> <frame-count> transition transparency <model-or-atom-spec> <start-transparency> <end-transparency> <frame-count> transition msms_transparency <msms-model-id> <start-transparency> <end-transparency> <frame-count> transition radial_motion <multiscale-surface-spec> <scale-factor> <frame-count> transition icosahedron_radius <start-radius> <end-radius> <radius-step>
Using ~transition reverses the direction of the transition, reversing the effect of the command without the "~" for explode and icosahedron commands but not for contour level or transparency adjustment.
Xing Zhang made radial fly away movies with other software. The radial motion command was written to make this easy to do in Chimera. The icosahedron radius transition was written for Tim Baker's lab that has made slicing movies with other tools (maybe ROBEM).
September 21, 2007. Updated for Chimera version 1.2430 where volume API changes were made to drop support for multi-component data.
February 27, 2007. Updated for Chimera version 1.2348 which uses NumPy instead of Numeric Python.
September 15, 2006. Added options to fade models in and out by varying transparency.
January 5, 2006. Initial version.