Tom Goddard
February 27, 2006
This Chimera tool displays steric clashes in crystal structures between different asymmetric units. It was written to validate crystal symmetry header information in Protein Data Bank virus structures.
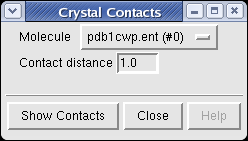
To show the crystal contacts dialog use menu entry
Tools / Higher-Order Structure / Crystal Contacts
Select the molecule, specify a contact distance, and press the Show Contacts button.
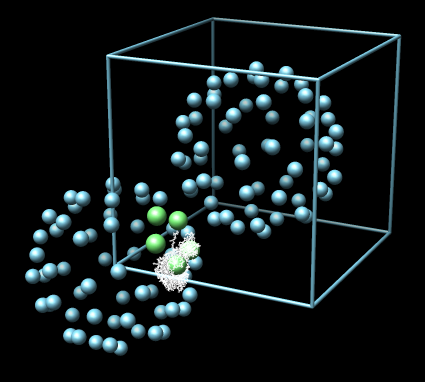
Pressing Show Contacts creates a new Chimera model where several small balls are shown, each ball representing one copy of the atomic coordinates contained in the PDB file. The different balls are positioned according to the crystal packing. Three ball colors are used. The collection of all green balls represents one crystal asymmetric unit. The collection of all green and blue balls represents one crystal unit cell. The yellow balls are from crystal asymmetric units in other unit cells.

|

|
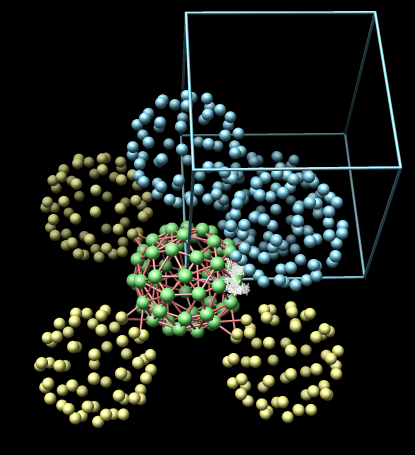
| Example 1: Cowpea Mosaic Virus (1ny7). Crystal symmetry information is correct with no 1A contacts. | Example 2: Cowpea Chlorotic Mottle Virus (1cwp). Crystal symmetry information is correct although there are a few atoms making 1A contacts between subunits. |
For many PDB models atomic coordinates are given for the entire crystal asymmetric unit. In these cases only one green ball will be shown. Virus capsid structures generally do not contain atomic coordinates for a full crystal asymmetric unit. They typically contain coordinates for one sixtieth (1/60) of one icosahedral virus particle because the icosahedron has 60-fold symmetry. Several copies (usually 5, 12, 15, 30, or 60) of the atomic coordinates are required to form the crystal asymmetric unit. We call one copy of the atomic coordinates the non-crystallographic symmetry (NCS) asymmetric unit. Each ball represents one NCS asymmetric unit. The MTRIX records in PDB file headers position copies of the atomic coordinates to create the crystal asymmetric unit.

|
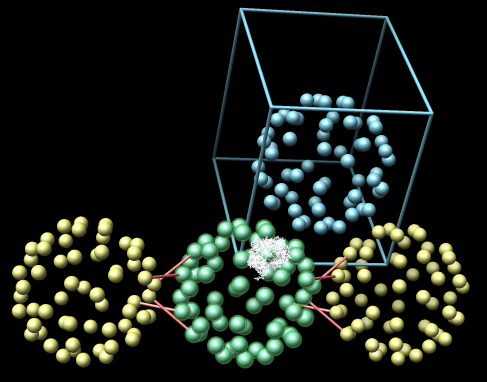
|
| Example 3: Conotoxin protein (1a0m) includes atomic coordinates for crystal asymmetric unit (one green ball). No 1A clashes. | Example 4: Rhinovirus 2 (1v9u). Transparent green balls covering blue balls indicates that one crystal asymmetric unit is directly on top of another. This is an error in crystal symmetry specification. |
Red cylinders connecting the balls indicate that the connected pair of NCS asymmetric units have contacting atoms. The radius of the cylinder is proportional to the number of contacting atoms. The minimum and maximum radii are limited with the minimum corresponding to 1% of all atoms in contact and maximum corresponding to 5%. For clarity cylinders are only shown that connect with the crystal asymmetric unit in green.
If two NCS asymmetric units (balls) are centered at the same point then the cylinder will not be visible since it is smaller than the ball. In this case the green ball is made partially transparent. The non-green balls are slightly smaller than the green ones and becomes visible inside. This condition indicates a definite error in the PDB symmetry specification.
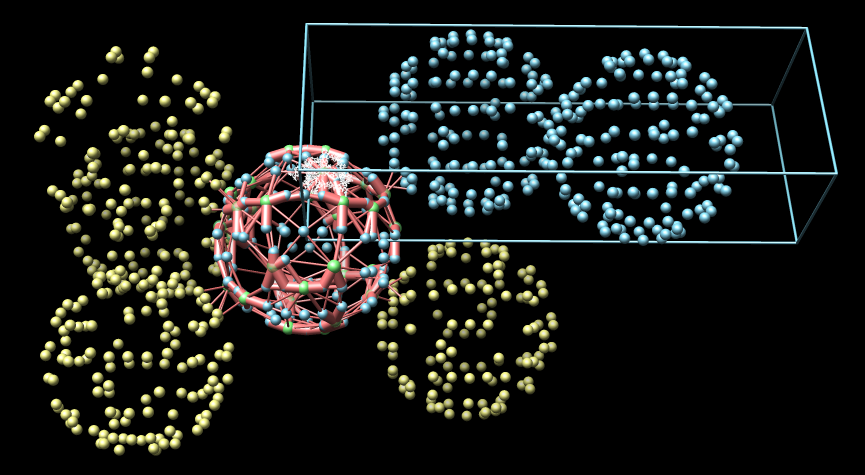
|
| Example 5: Bacteriophage MS2 (1bms). Lacks icosahedral symmetry and exhibits severe 1A clashes indicating a problem in symmetry specification. |
A blue box outlines one crystal unit cell. It has one corner at position (0,0,0).
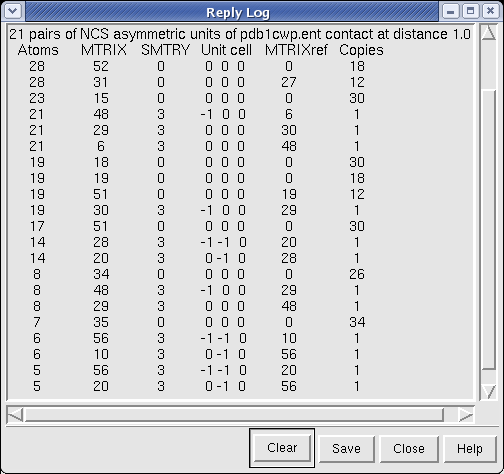
Pressing Show Contacts also displays a text table of contacting pairs of NCS asymmetric units in the Chimera Reply Log dialog. Each line reports the indices of matrices of a pair of contacting NCS asymmetric units. For each pair, and index MTRIXref tells which PDB MTRIX record positions one subunit, and indices (MTRIX, SMTRY, Unit Cell) identify the other subunit. The number of contact atoms in the first subunit is also reported. The MTRIX and SMTRY matrix indices start at 0, and therefore do not exactly match the matrix numbers given in the PDB header.
Each unique relative orientation of contacting NCS asymmetric units is reported, and the number of occurences of geometrically equivalent pairs with the same contacts is given in the Copies column of the table. Because of round-off errors in computing symmetry matrices, tolerances are used to identify equivalent relative orientations. These are 0.1 degrees for rotation and 0.1 Angstroms for translation.
There is a crystalcontacts Chimera command which shows a contacts model and contacts table just like the dialog interface. The command takes two arguments, the model number and the contact distance. For example,
crystalcontacts #0 1.0
February 27, 2007. Updated for Chimera version 1.2348 to use NumPy instead of Numeric Python.
August 8, 2006. Added crystalcontacts Chimera command. Added crystalimages.py script for generating contact images for a set of PDB files.
August 1, 2006. Original version based on discussions with Cathy Lawson.