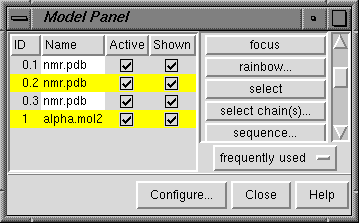
The Model Panel 
Each file of coordinates opened in Chimera becomes a
model with an associated model ID number.
Some PDB files are further subdivided into
multiple structures designated with MODEL and ENDMDL records;
thus, the
concept of submodels was created to allow unique atom specifications
even when multiple structures in the same file
contain identical residue and atom names.
Submodel numbers are assigned sequentially starting with 1 when the input
file contains more than one set of MODEL and ENDMDL
records (#0.1, #0.2, etc.); the numbers in the file are not used.
Each submodel can be treated as a separate model.
Thus, "models" will be used to indicate submodels and/or
models that are not subdivided into submodels.
Surfaces and other nonmolecular objects
are also assigned model ID numbers.
Molecules, surfaces, and objects can share a number.
The Model Panel shows the current models and conveniently enables
many operations upon them.
There are several ways to start
the Model Panel, a tool in the Inspectors and
Utilities categories.
Once one or more models have been chosen
within the left side of the Model Panel, any of several
functions represented by buttons on the
right side of the panel can be executed. Each
function is classified as frequently used
or infrequently used; the toggle button below the list
controls which set is listed.
Individual models or blocks of models can be
chosen (highlighted) with the left mouse button.
Ctrl-click adds to an existing choice rather than replacing it.
To highlight a block of models without having to hold down the mouse button,
click on the first (or last) and then Shift-click on the
last (or first) in the desired block.
In the figure, model 1 and submodel 2 of model 0 have been chosen.
Configure... allows
customization of which settings are shown as
checkboxes on the left side of the panel,
which functions are considered frequently used,
and
what functions are applied to double-clicked models.
Close dismisses the Model Panel. Help
brings up the Model Panel manual page in a browser window.
The functions are:
- activate
- activate for motion;
the opposite of deactivate
- activate all
- activate all models even if not chosen in the left side of the panel
(the previous pattern of activities is saved and can be restored using
the button's alternate function, restore activities)
- activate only
- activate the chosen model(s) and deactivate all the others
- attributes...
list model attributes and allow changes
- BLAST on PDB
- BLAST the sequence of a single model against sequences in the PDB using the
NCBI BLAST program blastcl3 (Network BLAST, see
BLAST Basic Overview).
This will only work for proteins, and the button will only appear
for users able to run blastcl3 from the system
command line (with arguments but no preceding path).
The E-value and matrix used can be adjusted.
Results are converted to a multiple sequence alignment and displayed
using MSF Viewer.
- clipping...
open the
Per-Model Clipping tool
- close
- close (remove) the model(s)
- compute SS
- define the secondary structure of peptides/proteins in the chosen model(s)
using ksdssp with the specified
parameter values. Hitting return (Enter) or clicking OK runs
the calculation and dismisses the dialog, while clicking Apply
runs the calculation without dismissing the dialog. Values saved as
defaults will only be used when peptide/protein structures that lack
HELIX and SHEET records are read in or when compute SS
is specifically invoked.
- deactivate
- deactivate for motion;
the opposite of activate
- focus
- center the display on the chosen model(s)
- hide
- disable display of the chosen model(s) (see
display hierarchy);
the opposite of show
- rainbow...
rainbow-color the chosen model(s), incrementing the color either
by residue or by chain. The color range is
defined by five color wells,
corresponding to starting color, three intermediate interpolation
colors, and ending color. When the increment is by residue, each
chain is rainbow-colored separately.
WATER and HET chains
are not affected.
- select
- select
chosen model(s) for subsequent operations or commands
- select chain(s)...
select
subsequently specified chains in the chosen model(s)
- sequence...
open the sequence panel
showing chain sequences
- show
- enable display of the chosen model(s) (see
display hierarchy);
the opposite of hide
- show all atoms
- show the chosen model(s) and display all atoms within them
- show only
- show the chosen model(s) and hide all of the others
- surface ligand
- display the molecular surface of atoms in the chosen model(s)
determined to be in the ligand category,
if any; equivalent to the typed command
surface ligand
- surface main
- display the molecular surface of atoms in the chosen model(s)
determined to be in the main category;
equivalent to the typed command
surface main
- tile
- "tile" the chosen models as described for
EnsembleTile; at least two models must be chosen for
this operation to be available
- toggle active
- toggle the status of each chosen model between
activated and deactivated for motion
(make the status of each its current opposite)
- trace backbones
- in each chosen model,
display only a simplified trace for bonded chains containing
amino acid or nucleic acid residues, consisting of real backbone bonds
(-[N-CA-C]n- in
peptides and -[O5'-C5'-C4'-C3'-O3'-P]n- in nucleic acids)
- trace chains
- in each chosen model,
display only a simplified trace for bonded chains containing
amino acid or nucleic acid residues, consisting of one atom per residue
(connect the CA atoms in peptides, C4' atoms in nucleic acids)
- transform as...
transform each (initially) chosen model using the rotation/translation matrix
of the single model chosen when the dialog's OK button is clicked
- write PDB
- open a dialog to
save models as PDB files. If multiple models are chosen, it is
possible to save them to a single PDB file (separated by END records) or
to multiple PDB files. Multiple PDB files are named by appending
model ID number (model or model.submodel)
to the specified file name, before the ".pdb" suffix, if any.
By default, the current transformed coordinates are saved.
Optionally, coordinates can be saved relative to the untransformed
coordinates of any model by checking the box marked Save relative
to model..., choosing the desired reference model in the
Model Panel, and then then clicking OK.
Model Panel Configuration
The Configure... button brings up another panel with three
sections for customizing the Model Panel. The three sections are
organized like index cards with their names on tabs across the top:
Buttons, Columns, and Double Click. Clicking on
a tab brings the corresponding card to the front.
-
The Buttons section
shows all the functions
that can be listed on the right side of the Model Panel.
If a checkbox is activated, the adjacent function
will be included in the frequently used list; the remaining
functions are included in the infrequently used list.
The frequently used/infrequently used toggle button is
below the function list on the right side of the Model Panel.
-
The Columns section
determines which model attributes will
be shown in the left side of the Model Panel:
- Active - whether the model is
activated for motion;
see activate
and deactivate
- ID - model (and submodel, if applicable) number
- Input file - pathname, relative if the file is in a subdirectory of
the working directory, otherwise a full pathname; if the file was
opened using a 4-character PDB identifier, only the PDB identifier is shown
- Name - filename
- Shown - whether the model is shown;
see hide
and show
Show model color behind model name places a panel of the
model-level color behind the model name.
The color of a model when opened is its model-level color.
-
The Double Click section
controls which
functions
will be executed upon
a model when the model is double-clicked in the left side of the Model
Panel. The default is attributes...;
however, there can be zero or many functions in the list for execution.
Clicking an entry in the Function menu
on the right causes it to appear in the Execution list on the left.
Clicking an entry in the Execution list removes it from that list.
The functions will be executed in the order listed.