PDB 1m4x
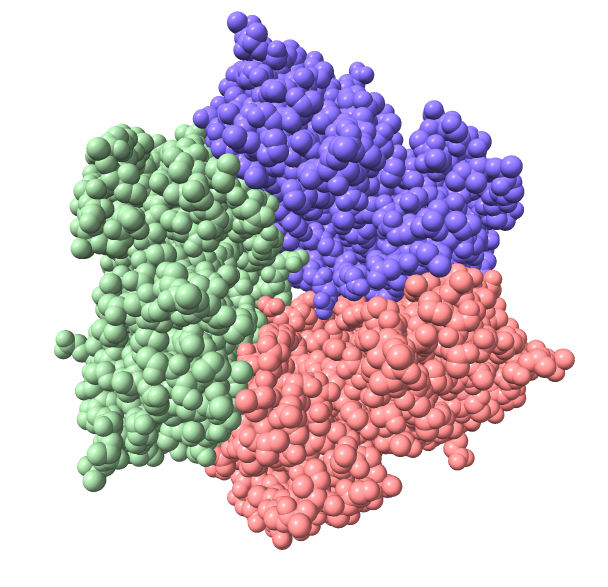
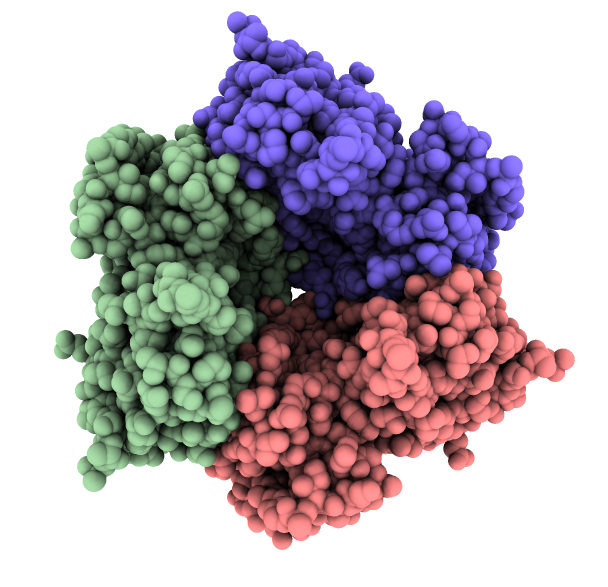
PDB 1aon
PDB 4v63
EMD 6035
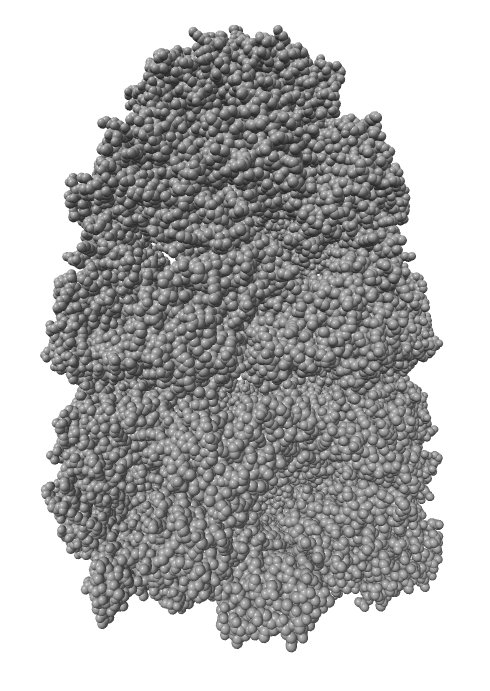
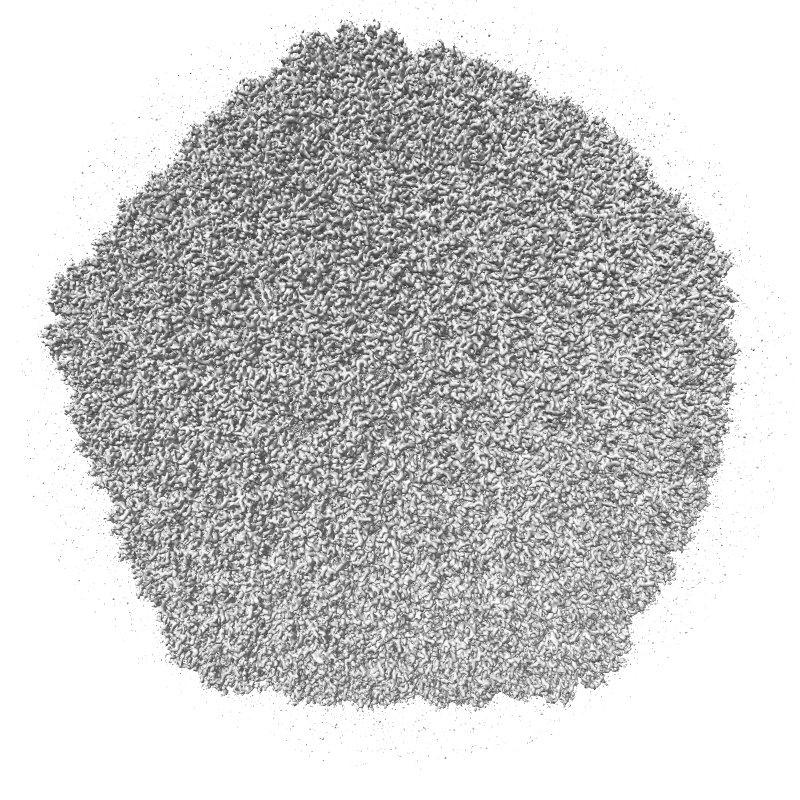

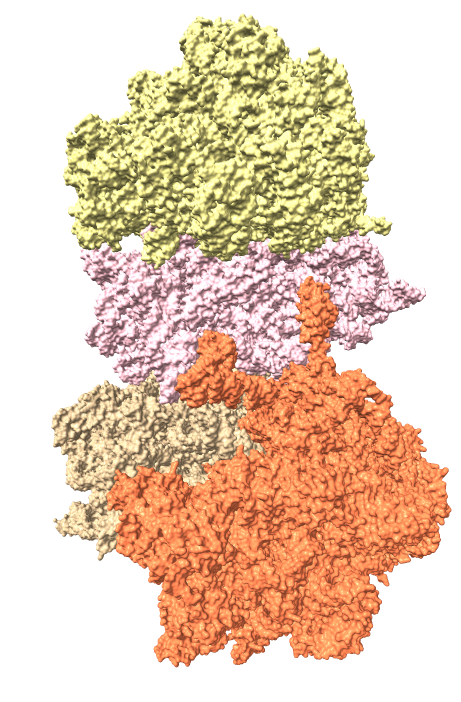

ambient shadows
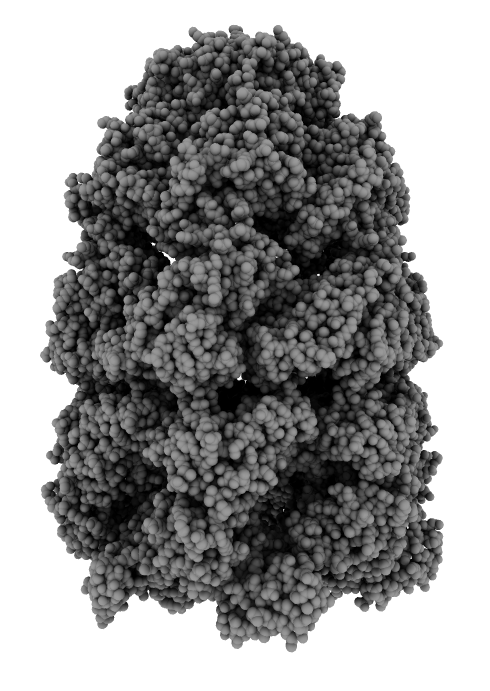
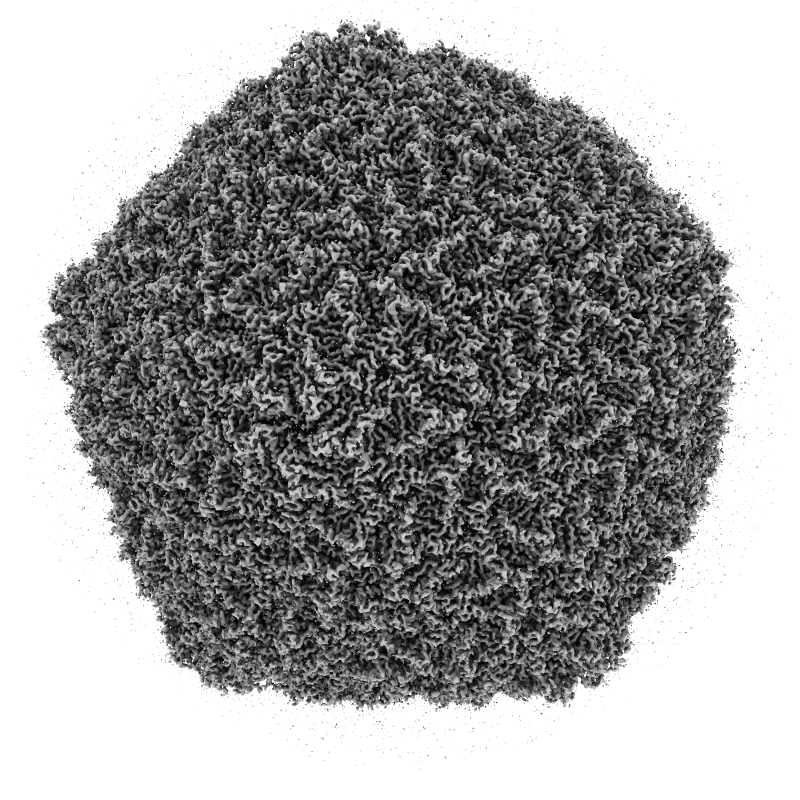


Advisory Committee
March 25, 2015
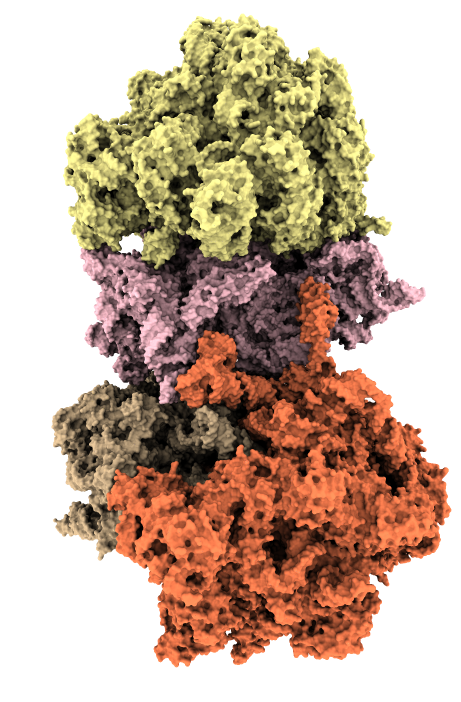
Ambient occlusion lighting. GPU programming allows interactive speed with 100 cast shadows.
| Virus capsomere PDB 1m4x | Chaperonin PDB 1aon | 70S ribosome PDB 4v63 | Bacteriophage T7 EMD 6035 | |
| Chimera 1 | 
| 
| 
| 
|
| ChimeraX ambient shadows | 
| 
| 
| 
|
Fast selection outlines.
|
Fast silhouette edges.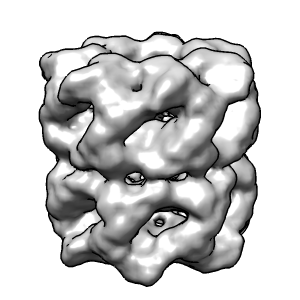
|
Fast molecule copies. 20,000 molecule HIV model. 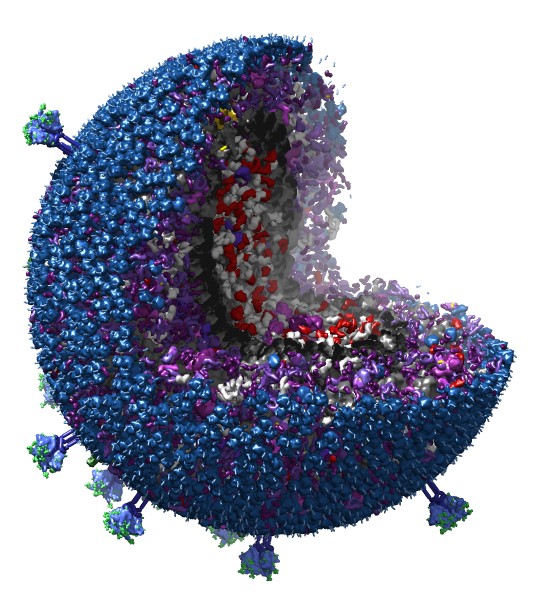
|
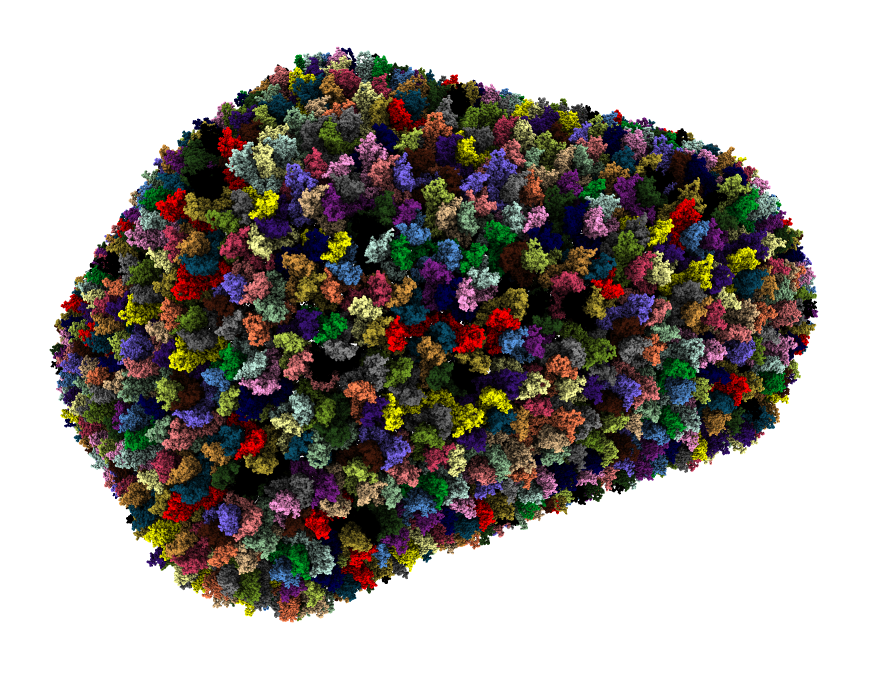

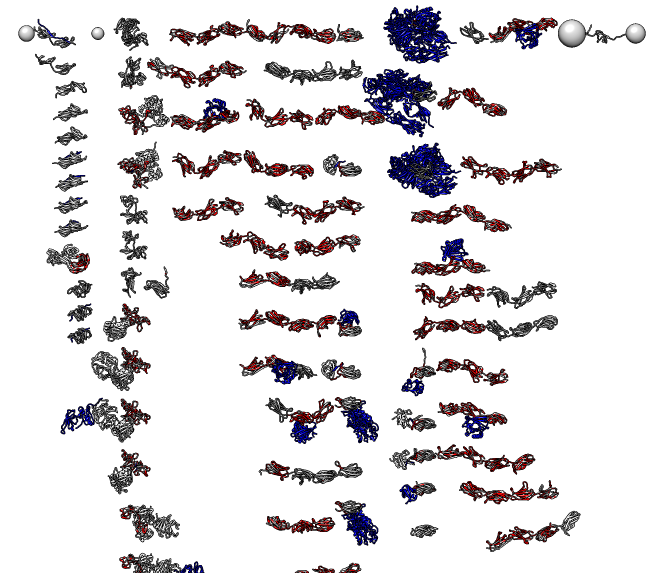
| 
|
| HIV capsid, 2.4 million atoms, 3j3q.cif | 106 PDB models matching fibronectin sequence P02751 |
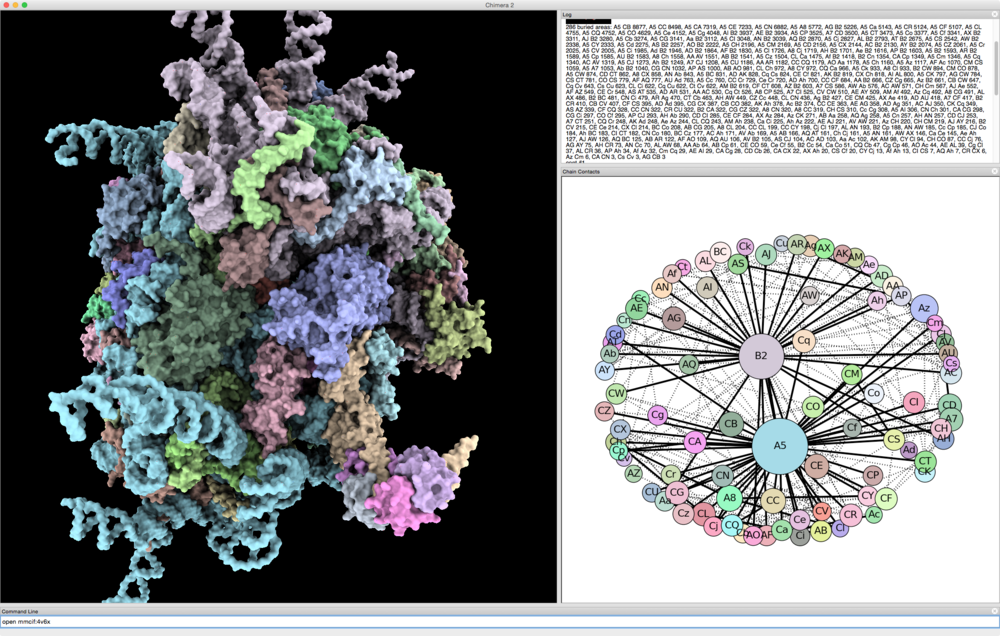
Some examples:

| 
| 
|
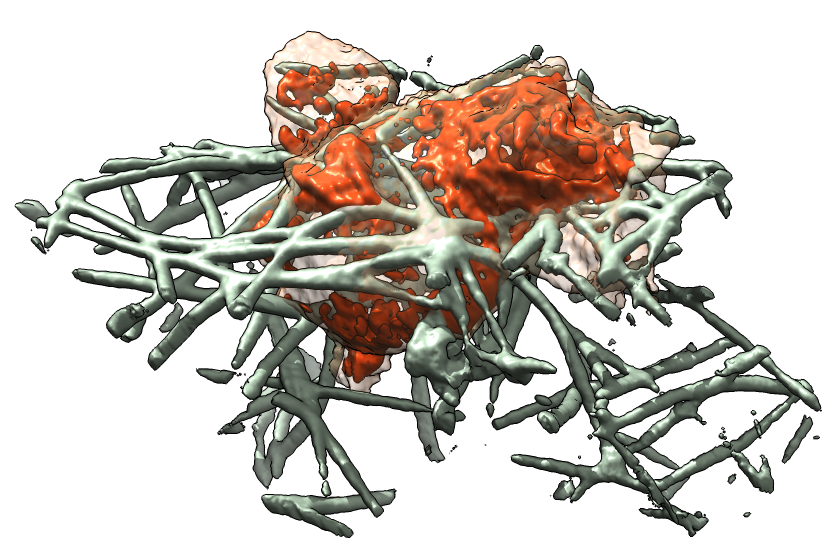
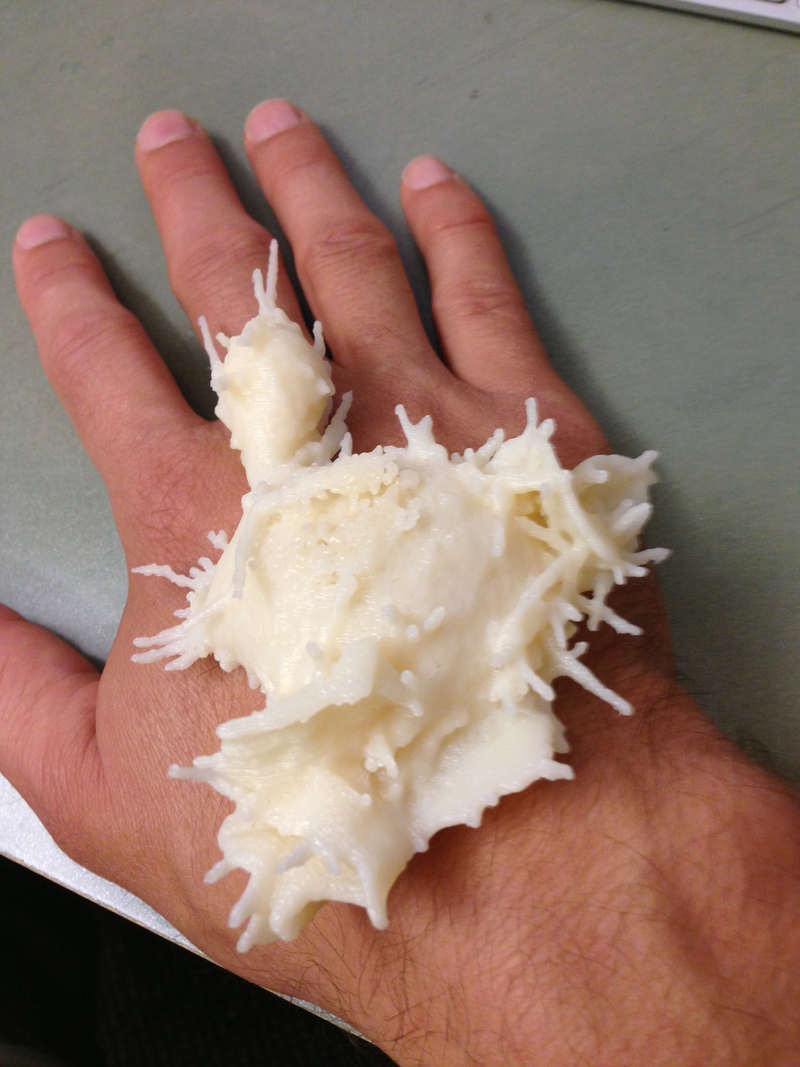
| Movie of cell motion. | Collagen only near cell path. | 3-D printed cell. |
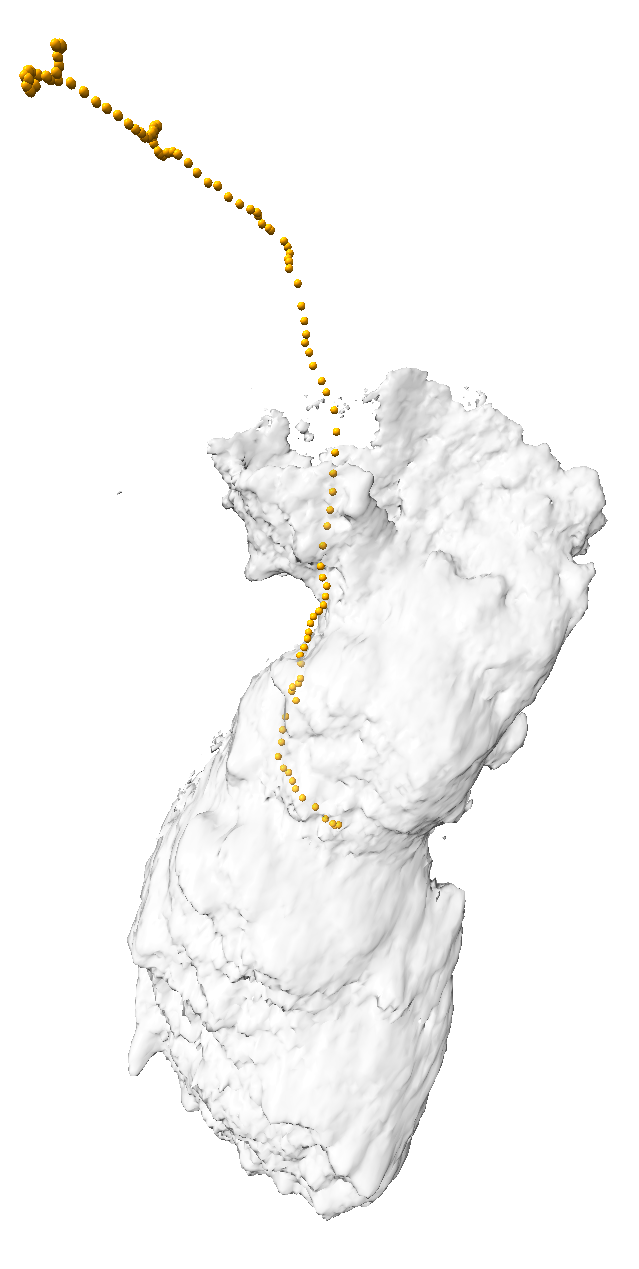
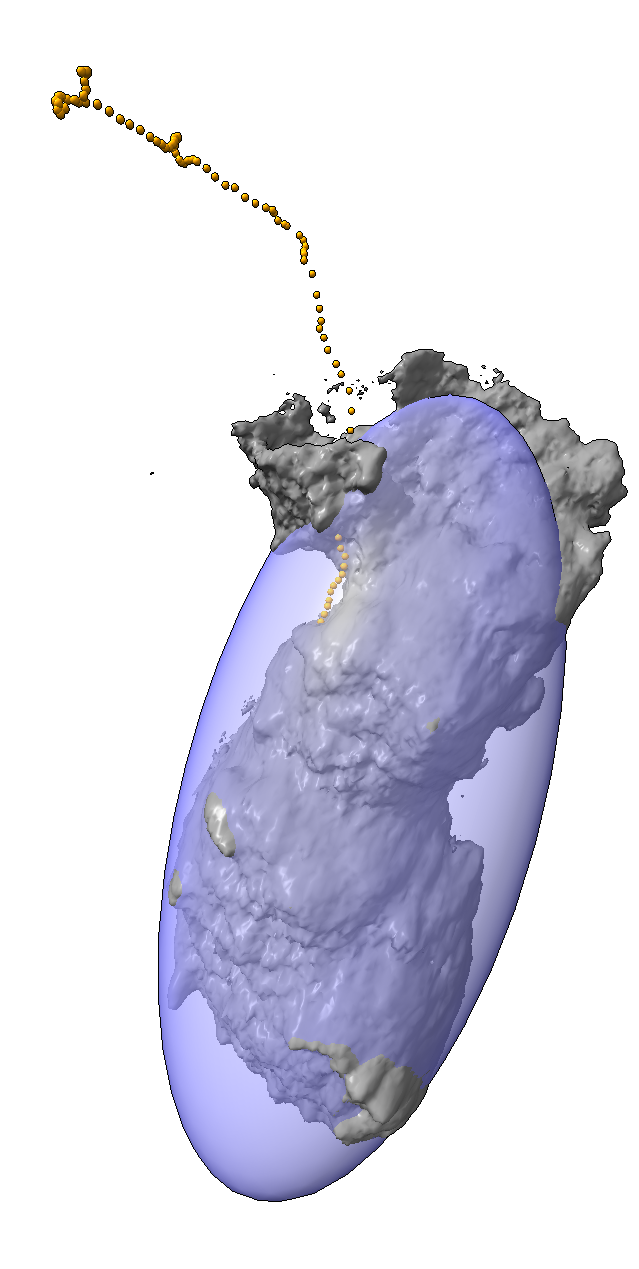
| 
| 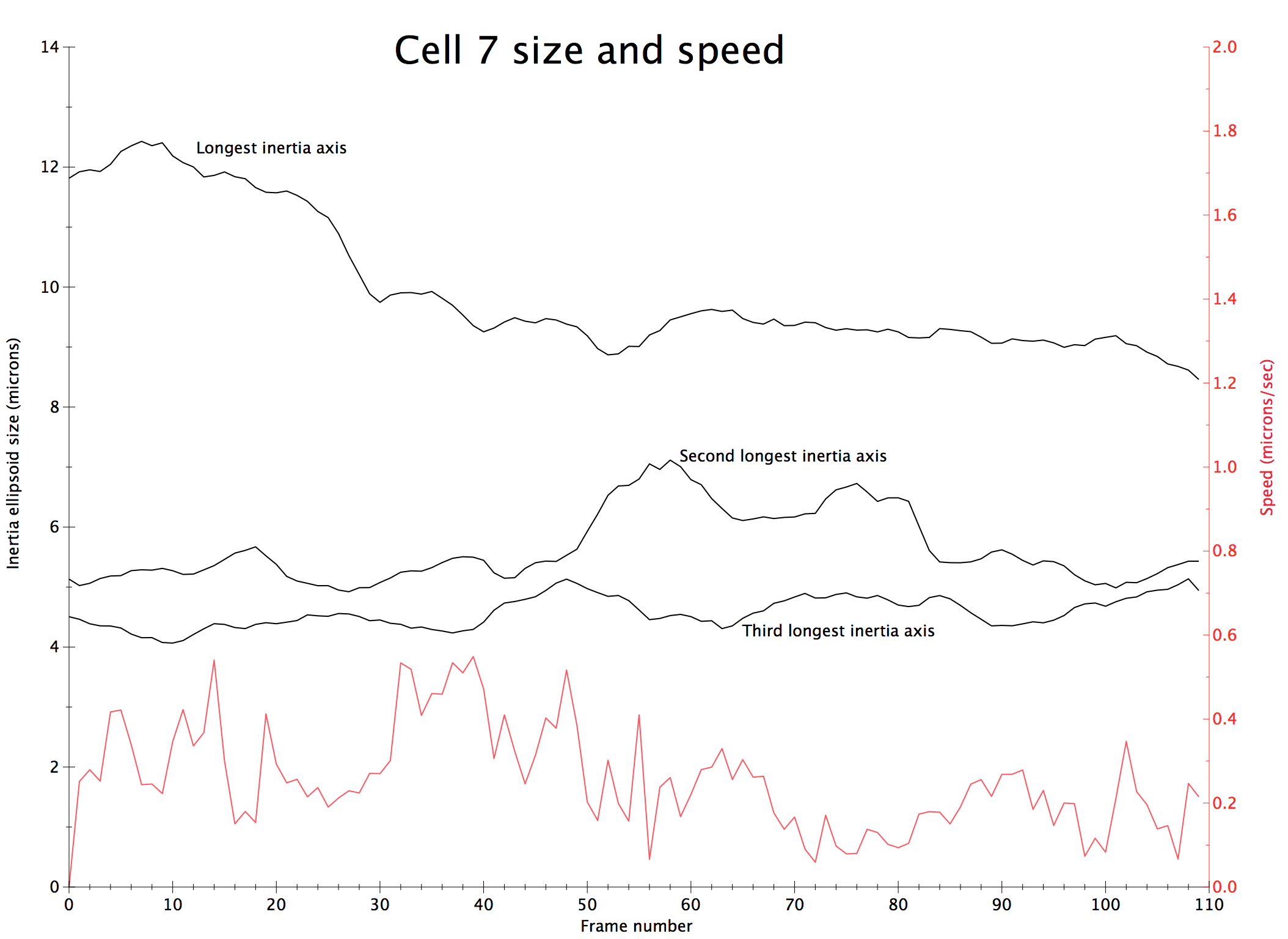
| 
|
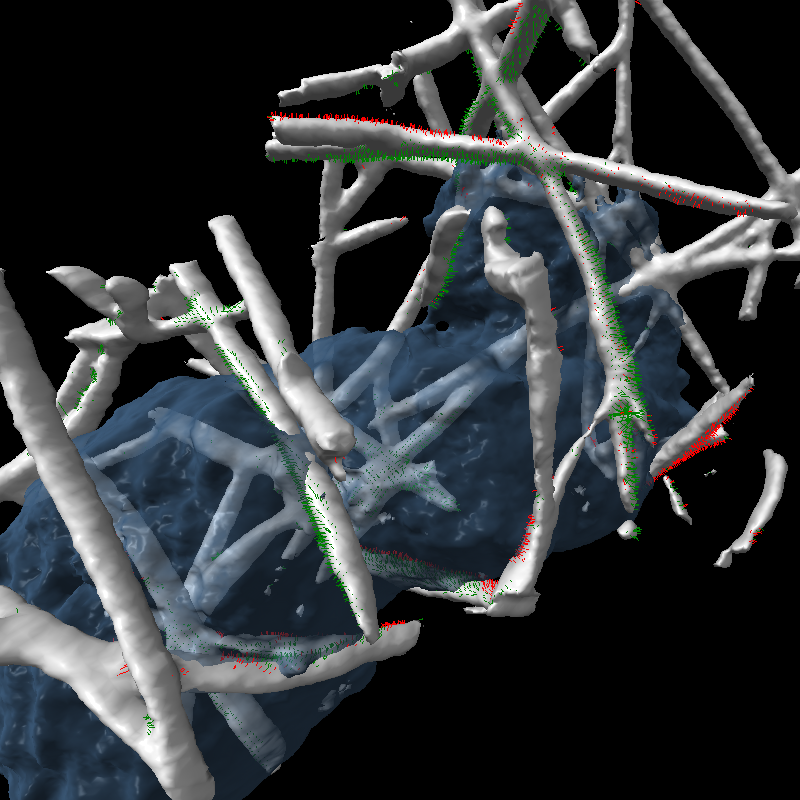
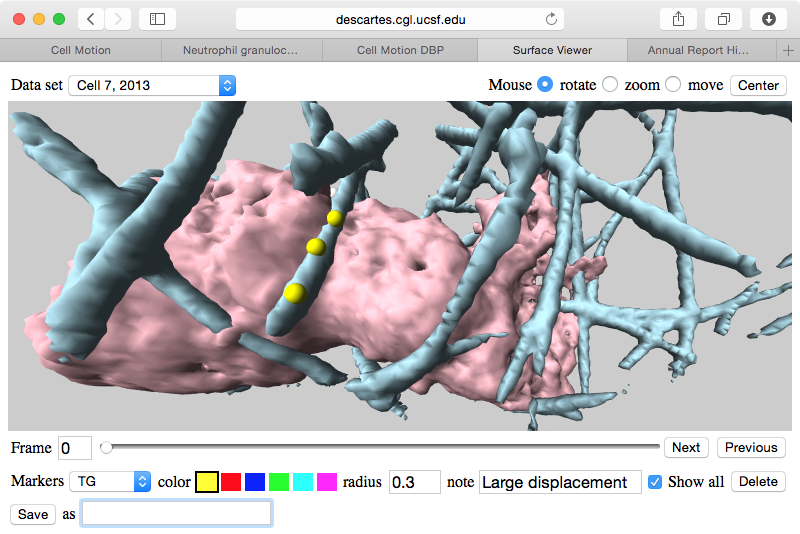
| Cell centroids (orange dots), size and axes (blue ellipsoid). | Plot cell speed and size changes. | Visualize filament motion (green and red prickles). | |
| 
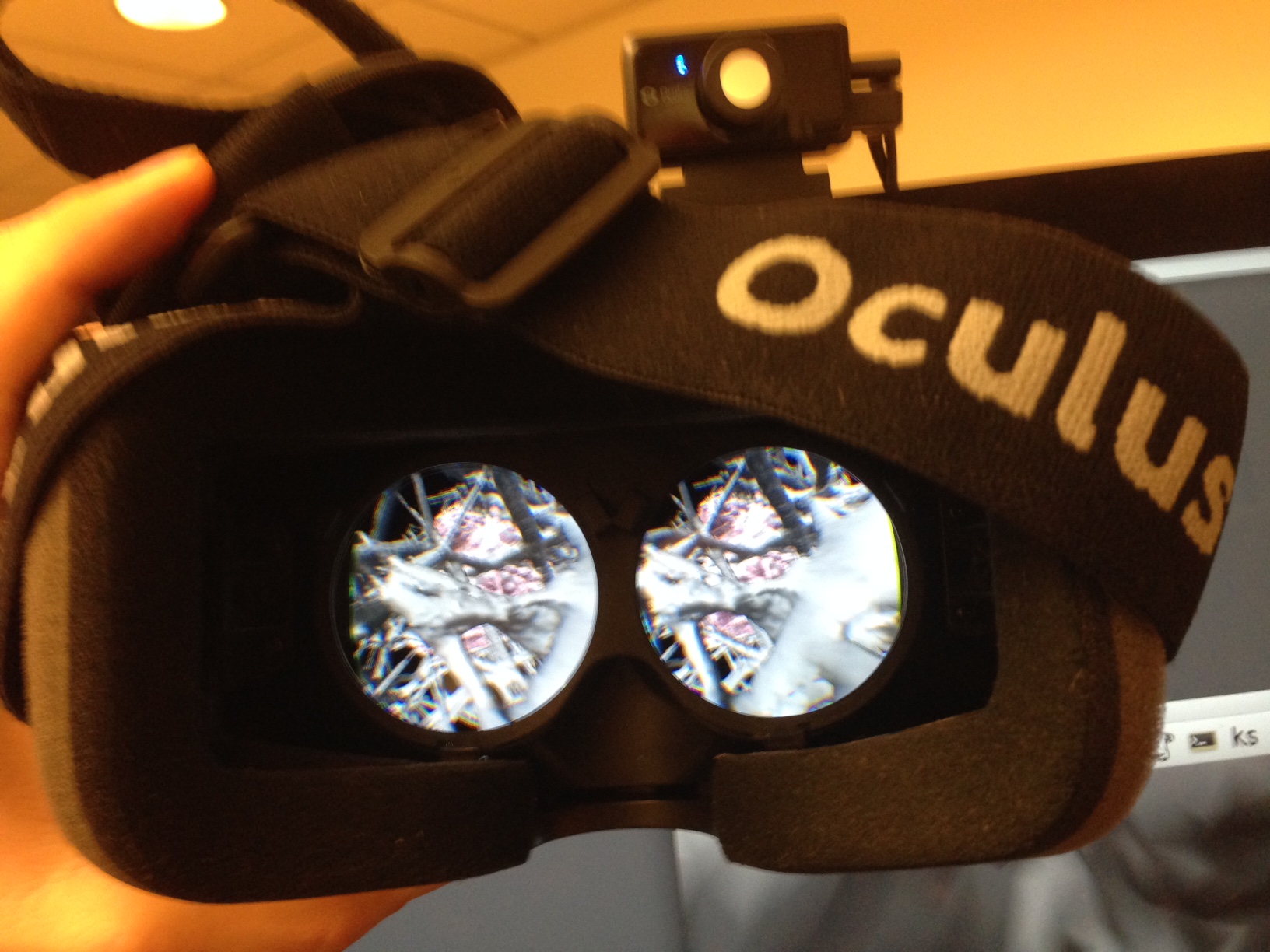
| 
|
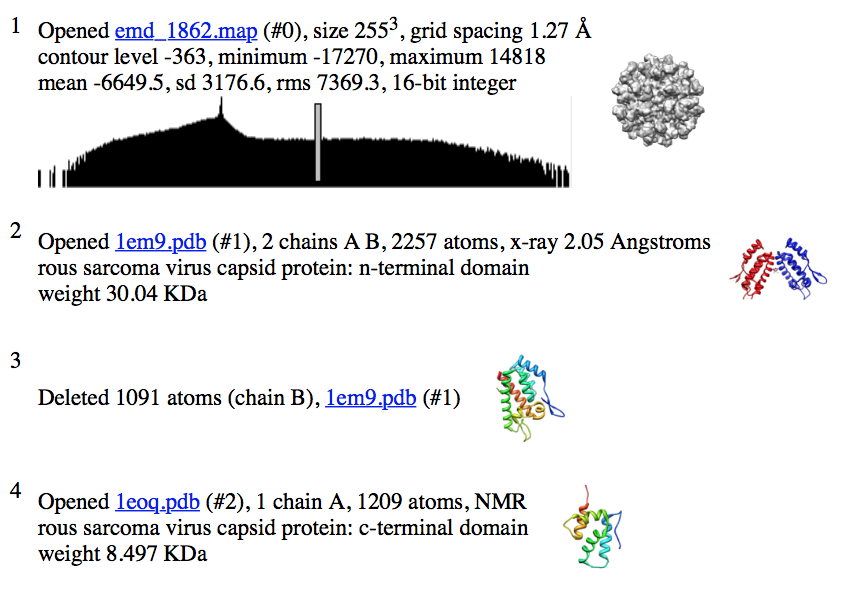

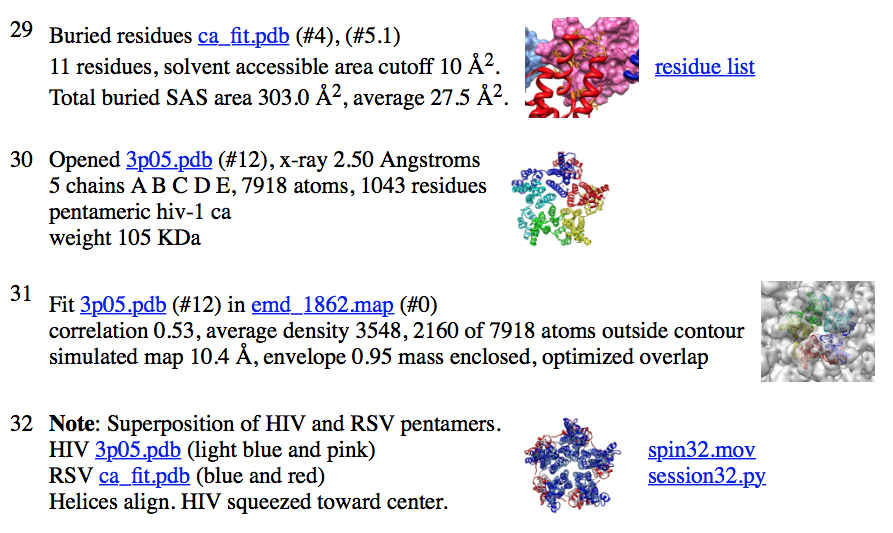
| ...
|

| 
| 
|
|
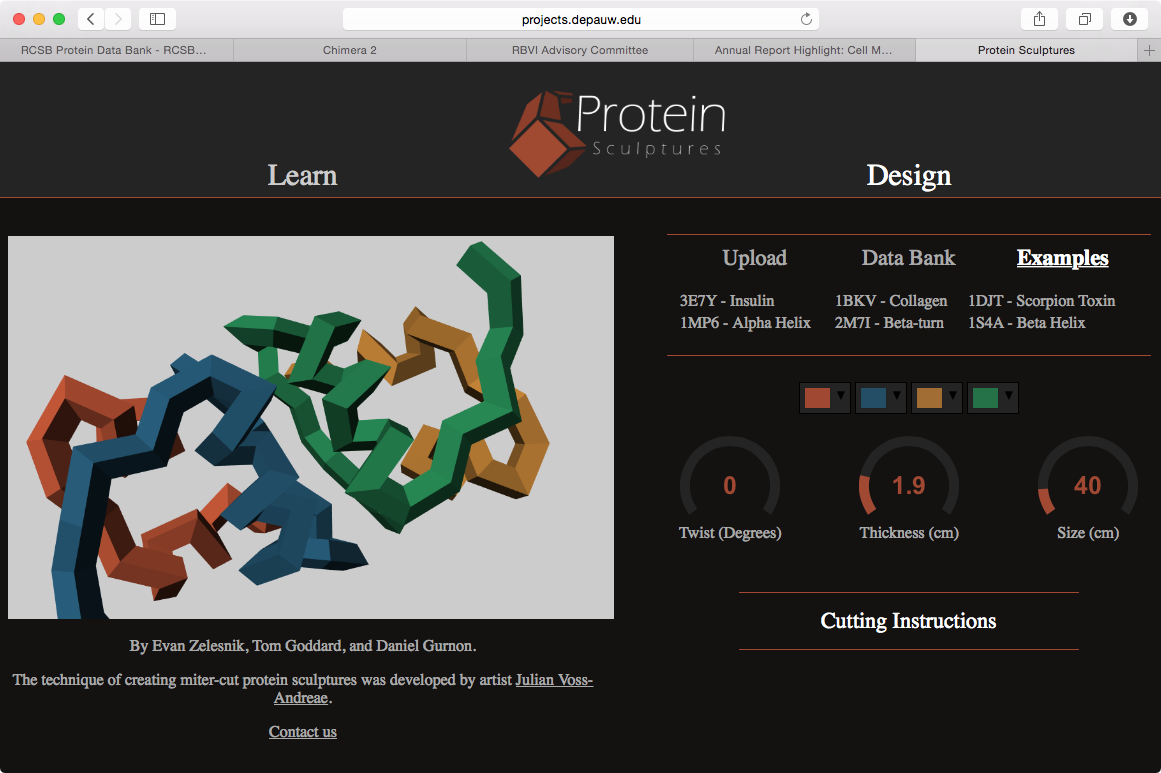
Alpha helix at Linus Pauling house. | Hemoglobin | Sculpture design Web app. |