
An ensemble contains multiple sets of coordinates for exactly the same atoms, for example, snapshots along a simulation trajectory or different solutions of NMR data.
Ensemble Match calculates superpositions and RMSD values for every pairwise comparison between two ensembles. Each ensemble must contain at least one structure (ensemble member), and the members of the two ensembles must contain the same set of atoms. Ensemble Match can also be used to compare an ensemble to itself. However, if the ensemble has many members, MD Movie may be more appropriate for obtaining all-by-all pairwise RMSD values (see RMSD analysis). See also: Ensemble Cluster, Tile Structures, superimposing structures
First, each ensemble should be opened in Chimera from a single PDB file (with MODEL and ENDMDL records delimiting individual members, if more than one). There are several ways to start Ensemble Match, a tool in the Structure Comparison and MD/Ensemble Analysis categories.
One ensemble should be designated as the Reference and the other as the Alternative. In the resulting table of comparisons, the reference structures will be listed vertically (the alternatives horizontally) and any matching operations will move the alternative structure onto the reference. Parts to Match are indicated with a command-line atom specification; if this field is left blank, all atoms will be used. The atom specification describes the atoms to be used in each structure. If atom names are given, they should specify equal numbers of atoms occurring in the same order in the different structures (if @ca is entered, the first CA in a structure is matched with the first CA in other structures, the second CA with the second in other structures, etc.).
OK performs the calculation and dismisses the dialog, while Apply performs the calculation without dismissing the dialog. Close dismisses the dialog without performing any calculation. Help brings up this manual page in a browser window.
|
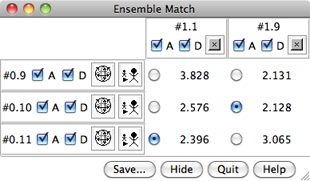
The resulting RMSD values are shown in a table. The figure at right shows comparisons between a reference ensemble with three members (models 0.8, 0.9, and 0.11) and an alternative ensemble with two members (models 1.1 and 1.9). In the table: