

 home
overview
research
resources
outreach & training
outreach & training
visitors center
visitors center
search
search
home
overview
research
resources
outreach & training
outreach & training
visitors center
visitors center
search
search


 home
overview
research
resources
outreach & training
outreach & training
visitors center
visitors center
search
search
home
overview
research
resources
outreach & training
outreach & training
visitors center
visitors center
search
search

Wah Chiu¹, and Tom Goddard²
¹
National Center for Macromolecular Imaging
Baylor College of Medicine
²
Resource for Biocomputing, Visualization, and Informatics
University of California, San Francisco
The National Center for Macromolecular Imaging (NCMI), run by Prof. Wah Chiu and located at the Baylor College of Medicine in Houston. The NCMI research has been devoted to developing electron cryomicroscopy (cryoEM) as a high resolution structural tool for biological assemblies that cannot be readily studied by X-ray crystallography or NMR spectroscopy. He has pioneered and demonstrated cryoEM reconstructions at subnanometer resolution (5-9Å) of several icosahedral viruses (Jiang et al., 2003; Zhou and Chiu, 2003; Zhou et al., 2000), GroEL (Ludtke et al., 2004), the calcium release channel (Ludtke et al., 2005) and the acrosomal bundle (Schmid et al., 2004). At these resolutions, not only can alpha helices and beta sheets can be identified, but protein folds can also be determined in some cases (Chiu et al., 2005; Jiang et al., 2001). The NCMI Resource Center has the mandate to collaborate with biologists from around the country, to provide access to its state-of-the-art technology, and to offer workshops on cryoEM technology. There are presently over 70 active collaborative projects. Generally, collaborative projects are initially carried out by the NCMI staff to generate sufficient preliminary results which then are used by either the collaborator or Dr. Chiu to seek independent grant support to continue a full investigation. Dr. Chiu's NIH R01 grant for studying structure and function of herpes simplex type 1 virus (R01AI38469, PI Chiu) and fatty acid synthase (R01GMS68826, PI Quiocho at Baylor College of Medicine) are two recent examples of successful collaborations initiated using NCMI resources.
All of the NCMI collaborative projects result in 3-D structures of relatively large biological complexes. The immediate task in interpreting the structure is to visualize the entire complex and to segment the individual subunits before the analysis of structural details such as helices and beta sheets. Steve Ludtke in our Center has adopted UCSF Chimera as one of the modules in the image processing pipeline within the EMAN software package (Ludtke et al., 1999). Among several locally-developed Chimera extensions, he has developed a module for producing animations using Chimera. These additions have enabled users to visualize the 3-D structures at any stage of the structural refinement. EMAN has over 300 users around the world. This software is freely accessible via the internet.
In a subproject of a Program Project (P01GM064692 directed by RM Glaeser at UC Berkeley), Chiu is responsible for collecting images of GroEL at the highest possible resolution using the liquid helium electron microscope at NCMI. Because the crystal structure of GroEL is known, this data set is intended to become a standard reference set and will be made accessible to the cryoEM community to validate development of novel image processing techniques. For instance, Steve Ludtke has recently used this data set to reconstruct a 4.5Å resolution map of GroEL using the latest improvements to his EMAN software suite (unpublished). The image below provides an example of the display of the final cryoEM maps of GroEL reconstructed at different resolutions (Ludtke et al., 2004; Ludtke et al., 2001).
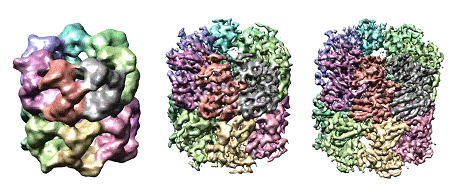
Figure caption: CryoEM maps of GroEL reconstructed using EMAN and imaged using Chimera at 11.5Å (left), 6Å (middle) and 4.5Å (right), with each of the 14 subunits colored differently.
Under NSF support (EIA-0325004), Chiu and his Co-PIs (Andrej Sali at UCSF and Chandrajit Bajaj at UT Austin) are developing new algorithms for structure visualization and interpretation of medium resolution structures determined by cryoEM. Chimera has become a critical module in the Analysis of Intermediate Resolution Structures (AIRS) software package that has been developed by Matthew Baker in our Center for structural analysis of cryoEM maps under this NSF support. The AIRS package includes twenty Chimera dialogs implemented by NCMI. Most of these dialogs setup calculations performed by command-line EMAN programs and then display results in the Chimera graphics window. Using Chimera for setting up input and analyzing output allows efficient interactive tuning of analysis parameters. Some of these tools identify probable locations of alpha helices and beta sheets in density maps, perform bilateral, median and other map filtering, perform fitting of flexible models in maps, and offer numerous segmentation methods for extracting substructures from maps. Cooperation between NCMI and RBVI enable development of these extensions and involves discussions of Chimera programming details and modifications to Chimera by the RBVI when needed.
In addition to the support that the RBVI provides enabling our Center to develop novel Chimera extensions, they have themselves developed several important enhancements to Chimera guided by our discussions. These include the capping and coloring the holes left when slicing density maps, optimizing the fit of models in cryoEM density maps, creating a 64-bit version for loading large data sets, and adding MRC 2000 file format support (handling data origins). Additional enhancements will be vital for the analysis of higher resolution maps. Interactive display and manipulation of large maps, gigabytes in size, is already a significant obstacle. More facile handling of the numerous segmented pieces of maps is also a high priority.
The collaboration with the RBVI extends beyond the typical P41 collaborative project. Also included are co-hosting of workshops. For example, on March 16-19, 2005, Dr. Ferrin and Mr. Tom Goddard traveled to Houston and taught in the NCMI workshop titled "Single Particle Reconstructions and Visualization: EMAN Tutorial", typically held biennially. Chimera was the primary visualization tool used at this workshop, and Dr. Ferrin gave a lecture describing Chimera's capabilities and Tom Goddard gave both a talk and a hands-on tutorial session on the use of Chimera. More than 100 people attended, several coming from Europe.
References:
- Chiu, W., Baker, M. L., Jiang, W., Dougherty, M., and Schmid, M. F. (2005). Electron cryomicroscopy of biological machines at subnanometer resolution. Structure (Camb) 13, 363-372.
- Jiang, W., Baker, M. L., Ludtke, S. J., and Chiu, W. (2001). Bridging the information gap: computational tools for intermediate resolution structure interpretation. J Mol Biol 308, 1033-1044.
- Jiang, W., Li, Z., Zhang, Z., Baker, M. L., Prevelige, P. E., and Chiu, W. (2003). Coat protein fold and maturation transition of bacteriophage P22 seen at sub-nanometer resolutions. Nat Struct Biol 10, 131-135.
- Ludtke, S. J., Baldwin, P. R., and Chiu, W. (1999). EMAN: Semi-automated software for high resolution single particle reconstructions. J Struct Biol 128, 82-97.
- Ludtke, S. J., Chen, D. H., Song, J. L., Chuang, D. T., and Chiu, W. (2004). Seeing GroEL at 6 Å resolution by single particle electron cryomicroscopy. Structure (Camb) 12, 1129-1136.
- Ludtke, S. J., Jakana, J., Song, J.-L., Chuang, D., and Chiu, W. (2001). A 11.5 Å single particle reconstruction of GroEL using EMAN. J Mol Biol 314, 253-262.
- Ludtke, S. J., Serysheva, II, Hamilton, S. L., and Chiu, W. (2005). The pore structure of the closed RyR1 channel. Structure (Camb) 13, 1203-1211.
- Schmid, M. F., Sherman, M. B., Matsudaira, P., and Chiu, W. (2004). Structure of the acrosomal bundle. Nature 431, 104-107.
- Zhou, Z. H., and Chiu, W. (2003). Determination of icosahedral virus structures by electron cryomicroscopy at subnanometer resolution. Adv Protein Chem 64, 93-124.
- Zhou, Z. H., Dougherty, M., Jakana, J., He, J., Rixon, F. J., and Chiu, W. (2000). Seeing the herpesvirus capsid at 8.5 Å. Science 288, 877-880.
Laboratory Overview | Research | Outreach & Training | Available Resources | Visitors Center | Search