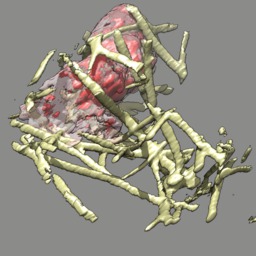
Tom Goddard
March 3, 2014
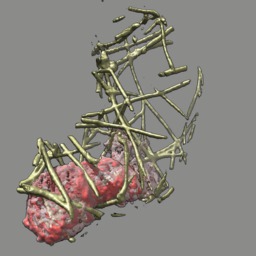
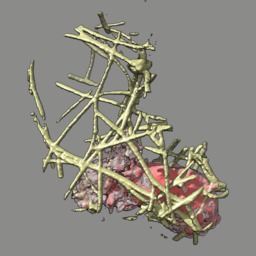
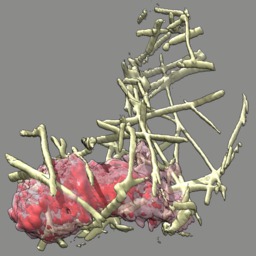
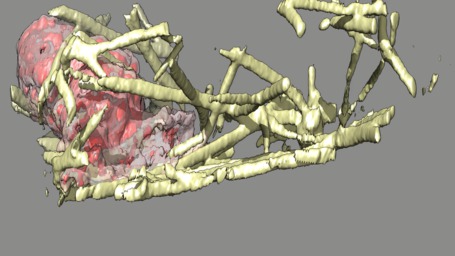
These videos of hl60 cells moving through collagen matrix show only the collagen that the cell touches at some point in the time course plus padding by 10 grid points (1 um in x,y and 2.5 in z). Data was acquired March 22, 2013 by Lillian Fritz-Laylin.
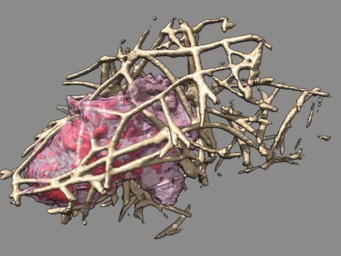
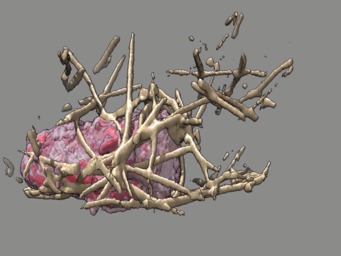
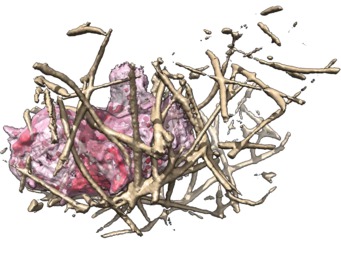
5 different view points. Click the image to play the movie.

| 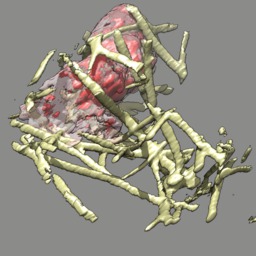
| 
| 
|
| above | approach | below | limbo |
|
| HD (1920 x 1080) |
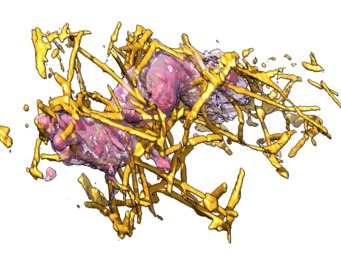
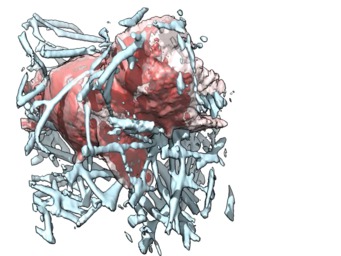
To show only the collagen touched by the cell first make a mask covering all the regions the cell visits, the pad that mask out a bit (1 micron), then apply that mask to the collagen maps. Here are example Chimera commands that do this. The saving of the cropped series took about 60 minutes per data set.
vseries open cell.cmap vop max # vop thresh #150 min 120 set 0 vop falloff #151 iterations 10 vop threshold #152 max 0 setMax 1 volume #153 save cell_mask.mrc vseries open collagen.cmap vseries save #154 collagen_masked.cmap mask #153 compress true vseries save #0 cell_masked.cmap mask #153 compress true
We compute a pointwise maximum ("vop max") of the cell maps at all times, set values below a specified threshold ("vop threshold") to zero, smooth the boundary of the mask to extend it by 10 grid points ("vop falloff"), then convert the mask to have 0 and 1 values ("vop threshold"). Finally we rewrite the collagen map series masking with this mask. Also we rewrite the cell map series with masking since this makes the series map file much smaller since compression is used so the parts with zeros take little space on disk.
Then to save a movie with the masked collagen.
vseries open collagen_masked.cmap vseries open cell_masked.cmap movie record supersample 3 ; vseries play #0 ; vseries play #150 ; wait 150 ; movie encode ~/Desktop/movie.mp4 quality high