- a UniProt accession number (example: P02751); UniProt's ID mapping service can be used to obtain UniProt accession numbers from other sequence database identifiers
- a pathname to a local file containing the sequence in FASTA format (example: /Users/miller/Desktop/mybpc3.fa); if the file contains multiple sequences, only the first will be used
 |
Performance depends on the number of structures imported into Chimera, which can be very large especially for runs with low filtering (permissive BLAST score and percent ID cutoffs, little or no winnowing) on queries with many known structures. Repeated uses of mda will reuse any open models that still meet the criteria and close any others. The command can be aborted during structure import by clicking the red stop icon in the status line to cancel the foreground task.
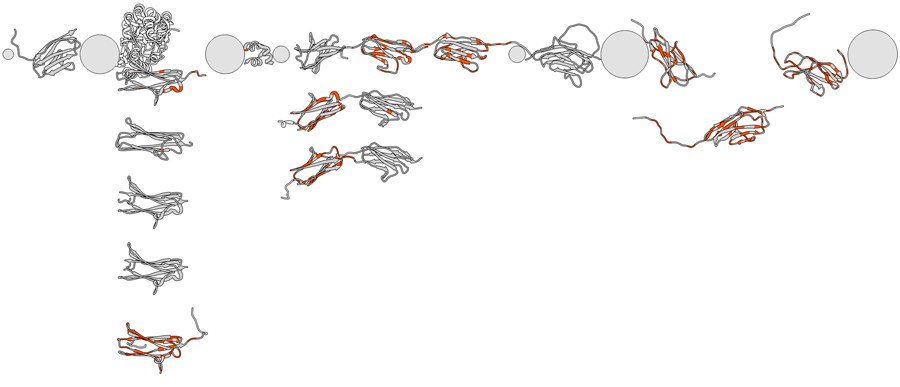
The center of rotation method is set to independent so that models rotate about their individual centers rather than a single collective center. The initial layout including scale and model orientations is saved as a position named stacked. Positions can be restored from the Rapid Access interface or with the command reset (e.g., reset stacked). Another position named overlay, better suited to the modeling step, is also saved.
- the pseudo-multiple sequence alignment from BLAST in aligned FASTA format
- a text file containing information from the run
On first use, mda also generates three database files (MDA_seqs.db, MDA_blast.db and MDA_uniprot.db) to speed up subsequent uses of the command, but these are not human-readable. These are placed in the user's Chimera download directory, in an MDA subdirectory.